
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 7,435,579 – 7,435,676 |
| Length | 97 |
| Max. P | 0.678769 |

| Location | 7,435,579 – 7,435,676 |
|---|---|
| Length | 97 |
| Sequences | 9 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 69.84 |
| Shannon entropy | 0.60663 |
| G+C content | 0.51134 |
| Mean single sequence MFE | -26.23 |
| Consensus MFE | -10.23 |
| Energy contribution | -9.81 |
| Covariance contribution | -0.42 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.678769 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 7435579 97 - 24543557 GCUCAGGCUCGGGAUGCCUAAUGCCCCGCAUUGUCUACGGCUUAAAGUCAACGACUC-UGUGGACUACAUUUCGCAGCCUCCUUGUUCGAGCAUUUCG------- (((((((((((..(((...(((((...)))))((((((((.....((((...)))))-)))))))..)))..)).)))))........))))......------- ( -32.60, z-score = -2.76, R) >droSim1.chr3L 6954285 97 - 22553184 GCCCAGGCUCGGGAUGCCUAAUGCCCCGCAUUGUCUACGGCUUAAAGUCAACGACUC-UGUGGACUACAUUCCGCAGCCUCCUUGUUCGAGGAUUUCG------- .....(((((((((((...(((((...)))))((((((((.....((((...)))))-)))))))..))))))).))))(((((....))))).....------- ( -35.00, z-score = -3.07, R) >droSec1.super_2 7366908 97 - 7591821 GCCCAGGCUCGGGAUGCCUAAUGCCCCGCAUUGUCUACGGCUUAAAGUCAACGACUC-UGUGGACUACAUUCCGCAGCCUCCUUGUUCGAGGAUUUCG------- .....(((((((((((...(((((...)))))((((((((.....((((...)))))-)))))))..))))))).))))(((((....))))).....------- ( -35.00, z-score = -3.07, R) >droYak2.chr3L 8040183 104 - 24197627 GGCCAGGCUCAGGAUGCCUAAUGCCCCGCAUUGUCUACGGCUUAAAGUCAACGACUC-UGUGGACUACAUUUCGCUGCCUGCCUAGUUGAGUAUUUCGCAUUUCG (((.((((.(..((((...(((((...)))))((((((((.....((((...)))))-)))))))..))))..)..)))))))...................... ( -28.00, z-score = -0.35, R) >droEre2.scaffold_4784 21856512 96 + 25762168 GGCCAGGCUCGGGCUGCCUAAUGCCCCGCAUUGUCUGCGGCUUAAAGUCAACGACUC-UGUGGACUACAUUUCGCAGGCUCCUUGGUCGAGUAUUCG-------- (((((((...((((........))))......((((((((.....((((.(((....-))).)))).....))))))))..))))))).........-------- ( -34.30, z-score = -1.52, R) >droAna3.scaffold_13337 16225316 81 - 23293914 -----------GAAAGCCUAAUUCCUCGUAUUGUCUACGGCUUAAAGUCAACGACUU-UGUGGACUACUUUCCCCCGUCAUUUUGUUAUUUUU------------ -----------(((((......((((((((.....)))))..(((((((...)))))-)).)))...))))).....................------------ ( -11.10, z-score = -0.13, R) >dp4.chrXR_group6 1432726 76 - 13314419 GUCCGGGAUCGAGCCGCCUAAUGCCCUGCAUUGUCUACGGCUUAAAGUCAACGACUUGUGUGGACUAUAUUUCUGG----------------------------- ..((((((..((((((..((((((...))))))....))))))..((((.(((.....))).))))....))))))----------------------------- ( -21.90, z-score = -1.41, R) >droPer1.super_12 2122420 76 - 2414086 GUCCGGGAUCGAGCCGCCUAAUGCCCUGCAUUGUCUACGGCUUAAAGUCAACGACUUGUGUGGACUAUAUUUCUGG----------------------------- ..((((((..((((((..((((((...))))))....))))))..((((.(((.....))).))))....))))))----------------------------- ( -21.90, z-score = -1.41, R) >droWil1.scaffold_180698 5963063 82 - 11422946 --GCUGCUGCAGGC--CUUAAUACGCUGCAUUGUCUACGGCUUAAAGUCAACGACUU-UGUGGACUAUUUCCCGGUUAAUUUUAUGU------------------ --((....)).(((--(..((((..(..((..(((...(((.....)))...)))..-))..)..))))....))))..........------------------ ( -16.30, z-score = 0.31, R) >consensus GCCCAGGCUCGGGAUGCCUAAUGCCCCGCAUUGUCUACGGCUUAAAGUCAACGACUC_UGUGGACUACAUUUCGCAGCCUCCUUGUUCGAG_AUU__________ .........................(((((..(((...(((.....)))...)))...))))).......................................... (-10.23 = -9.81 + -0.42)
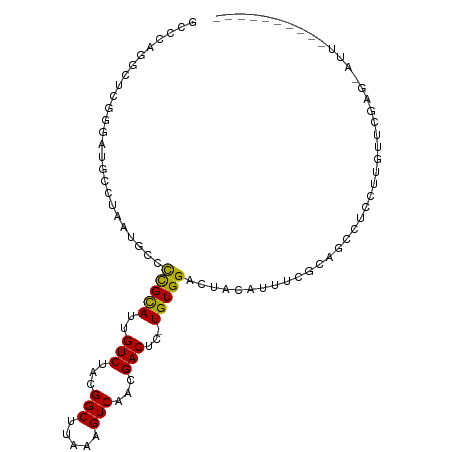


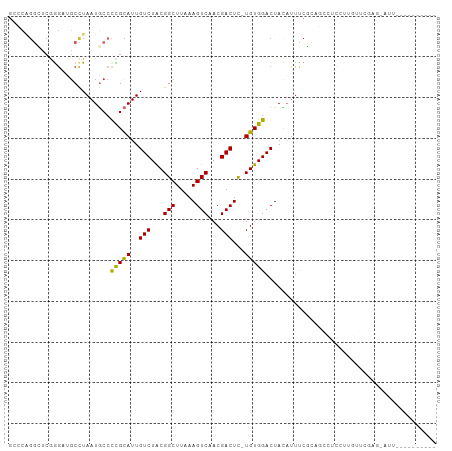
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:09:31 2011