
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 7,328,330 – 7,328,424 |
| Length | 94 |
| Max. P | 0.990711 |

| Location | 7,328,330 – 7,328,424 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 68.67 |
| Shannon entropy | 0.59396 |
| G+C content | 0.41494 |
| Mean single sequence MFE | -29.05 |
| Consensus MFE | -10.79 |
| Energy contribution | -10.68 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.44 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.43 |
| SVM RNA-class probability | 0.990711 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
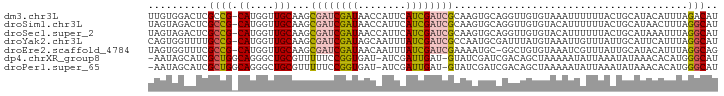
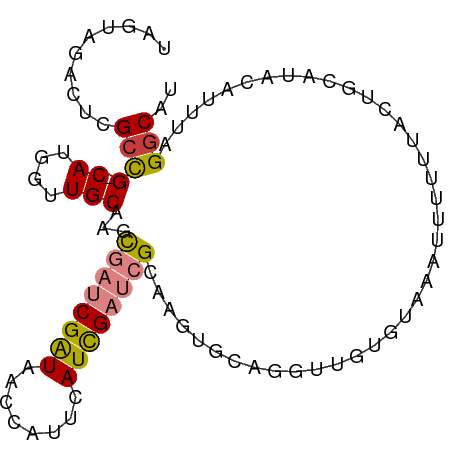
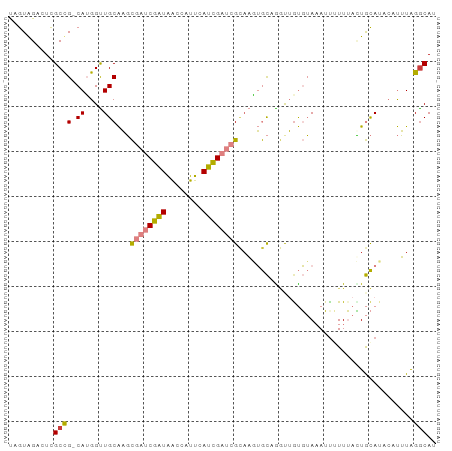
>dm3.chr3L 7328330 94 + 24543557 UUGUGGACUCGCCG-CAUGGUUGCAAGCGAUCGAUAACCAUUCAUCGAUCGCAAGUGCAGGUUGUGUAAAUUUUUUACUGCAUACAUUUAGACAU .(((((.....)))-))..(((....(((((((((........)))))))))..((((((.................)))))).......))).. ( -27.53, z-score = -1.93, R) >droSim1.chr3L 6845517 94 + 22553184 UAGUAGACUCGCCG-CAUGGUUGCAAGCGAUCGAUAACCAUUCAUCGAUCGCAAGUGCAGGUUGUGUACAUUUUUUACUGCAUAACUUUAGGCAU ..........(((.-......((((.(((((((((........)))))))))...))))(((((((((..........)))))))))...))).. ( -30.90, z-score = -3.24, R) >droSec1.super_2 7253169 94 + 7591821 UAGUAGACUCGCCG-CAUGGUUGCAAGCGAUCGAUAACCAUUCAUCGAUCGCAAGUGCAGGUUGUGUACAUUUUUUACUGCAUAAAUUUAGGCAU ..........((((-((....)))..(((((((((........)))))))))..((((((.................)))))).......))).. ( -29.73, z-score = -2.99, R) >droYak2.chr3L 7932921 94 + 24197627 CAGUGGUUUUGCCG-CAUGGUUGCAAGCGAUCGAUAGCAAUUUAUCGAUCGCCAAUGCGAUUUAUGUAAAUUGUUUAUUGCAUUCAUUUAGGCAU .........((((.-.(((..((((((((((((((((....)))))))))))....(((((((....)))))))...)))))..)))...)))). ( -32.70, z-score = -3.55, R) >droEre2.scaffold_4784 21748692 93 - 25762168 UAGUGGUUUCGCCG-CAUGGUUGCAAGCGAUCGAUAACAAUUUAUCGAUCGAAAAUGC-GGCUGUGUAAAUCGUUUAUUGCAUACAUUUAGGCAG ..........((((-((((((((((..((((((((((....))))))))))....)))-)))))))).....((.....)).........))).. ( -32.30, z-score = -3.53, R) >dp4.chrXR_group8 3419630 92 + 9212921 -AAUAGCAUCGCUGGCAGGGCUGCGUUUUUCCGGUGAU-AUCGAUUGAU-GUAUCGAUCGACAGCUAAAAAUAUUAAAUAUAAACACAUGGGCAU -..((((((((((((.(((((...))))).))))))))-.((((((((.-...))))))))..))))............................ ( -25.10, z-score = -1.95, R) >droPer1.super_65 223823 92 + 397436 -AAUAGCAUCGCUGGCAGGGCUGCGUUUUUCCGGUGAU-AUCGAUUGAU-GUAUCGAUCGACAGCUAAAAAUAUUAAAUAUAAACACAUGGGCAU -..((((((((((((.(((((...))))).))))))))-.((((((((.-...))))))))..))))............................ ( -25.10, z-score = -1.95, R) >consensus UAGUAGACUCGCCG_CAUGGUUGCAAGCGAUCGAUAACCAUUCAUCGAUCGCAAGUGCAGGUUGUGUAAAUUUUUUACUGCAUACAUUUAGGCAU ..........(((........((((...(((((((........)))))))((....))....................))))........))).. (-10.79 = -10.68 + -0.11)
| Location | 7,328,330 – 7,328,424 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 68.67 |
| Shannon entropy | 0.59396 |
| G+C content | 0.41494 |
| Mean single sequence MFE | -23.10 |
| Consensus MFE | -8.44 |
| Energy contribution | -9.50 |
| Covariance contribution | 1.07 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.823866 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
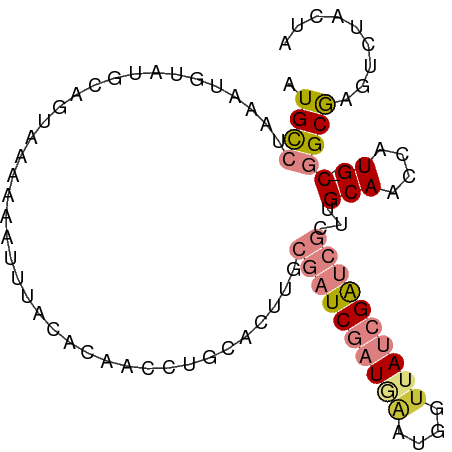
>dm3.chr3L 7328330 94 - 24543557 AUGUCUAAAUGUAUGCAGUAAAAAAUUUACACAACCUGCACUUGCGAUCGAUGAAUGGUUAUCGAUCGCUUGCAACCAUG-CGGCGAGUCCACAA ..(.((...((((((..((((.....))))......((((...(((((((((((....))))))))))).))))..))))-))...)).)..... ( -27.50, z-score = -2.50, R) >droSim1.chr3L 6845517 94 - 22553184 AUGCCUAAAGUUAUGCAGUAAAAAAUGUACACAACCUGCACUUGCGAUCGAUGAAUGGUUAUCGAUCGCUUGCAACCAUG-CGGCGAGUCUACUA .((((........((((........)))).......((((...(((((((((((....))))))))))).))))......-.))))......... ( -26.90, z-score = -2.17, R) >droSec1.super_2 7253169 94 - 7591821 AUGCCUAAAUUUAUGCAGUAAAAAAUGUACACAACCUGCACUUGCGAUCGAUGAAUGGUUAUCGAUCGCUUGCAACCAUG-CGGCGAGUCUACUA .((((........((((........)))).......((((...(((((((((((....))))))))))).))))......-.))))......... ( -26.90, z-score = -2.61, R) >droYak2.chr3L 7932921 94 - 24197627 AUGCCUAAAUGAAUGCAAUAAACAAUUUACAUAAAUCGCAUUGGCGAUCGAUAAAUUGCUAUCGAUCGCUUGCAACCAUG-CGGCAAAACCACUG .((((...(((..(((((....((((.............))))((((((((((......)))))))))))))))..))).-.))))......... ( -27.42, z-score = -4.18, R) >droEre2.scaffold_4784 21748692 93 + 25762168 CUGCCUAAAUGUAUGCAAUAAACGAUUUACACAGCC-GCAUUUUCGAUCGAUAAAUUGUUAUCGAUCGCUUGCAACCAUG-CGGCGAAACCACUA .(((.((....)).)))................(((-((((....(((((((((....)))))))))(....)....)))-)))).......... ( -23.20, z-score = -2.39, R) >dp4.chrXR_group8 3419630 92 - 9212921 AUGCCCAUGUGUUUAUAUUUAAUAUUUUUAGCUGUCGAUCGAUAC-AUCAAUCGAU-AUCACCGGAAAAACGCAGCCCUGCCAGCGAUGCUAUU- ............................((((.((((((((((..-....))))))-..............(((....)))...))))))))..- ( -14.90, z-score = 0.42, R) >droPer1.super_65 223823 92 - 397436 AUGCCCAUGUGUUUAUAUUUAAUAUUUUUAGCUGUCGAUCGAUAC-AUCAAUCGAU-AUCACCGGAAAAACGCAGCCCUGCCAGCGAUGCUAUU- ............................((((.((((((((((..-....))))))-..............(((....)))...))))))))..- ( -14.90, z-score = 0.42, R) >consensus AUGCCUAAAUGUAUGCAGUAAAAAAUUUACACAACCUGCACUUGCGAUCGAUGAAUGGUUAUCGAUCGCUUGCAACCAUG_CGGCGAGUCUACUA .((((........(((((...................((....))(((((((((....)))))))))..)))))........))))......... ( -8.44 = -9.50 + 1.07)

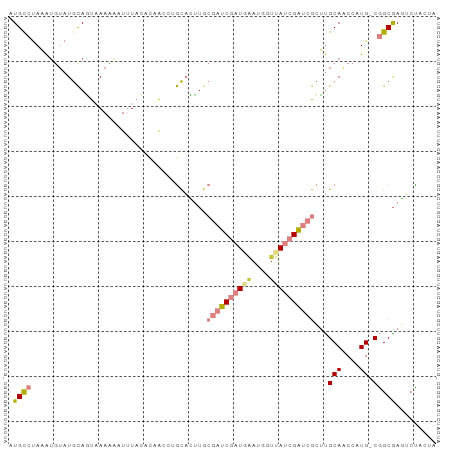
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:09:24 2011