| Sequence ID | dm3.chr3L |
|---|---|
| Location | 7,294,341 – 7,294,437 |
| Length | 96 |
| Max. P | 0.933021 |

| Location | 7,294,341 – 7,294,437 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 70.22 |
| Shannon entropy | 0.60576 |
| G+C content | 0.53264 |
| Mean single sequence MFE | -31.46 |
| Consensus MFE | -16.41 |
| Energy contribution | -17.05 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.933021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
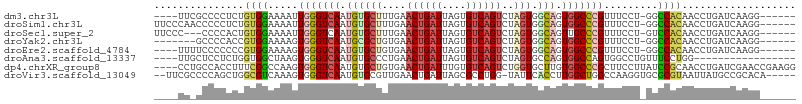
>dm3.chr3L 7294341 96 + 24543557 ----UUCGCCCCUCUGUGGAAAAUUGGGUCAAUGUGCUUUGAACUGAUUAGUGUCAGUCUAGUGGCAGUGGCCCGUUUCCU-GGCCACAACCUGAUCAAGG------ ----...(((.......(((((...((((((...(((((((.((((((....)))))).))).)))).)))))).))))).-))).....(((.....)))------ ( -29.40, z-score = -1.03, R) >droSim1.chr3L 6805997 100 + 22553184 UUCCCAACCCCCUCUGUGGAAAAUUGGGUCAAUGUGCUUUGAACUGAUUAGUGUCAGUCUAGUGGCAGUGGCCCGUUUCCU-GGCCACAACCUGAUCAAGG------ ((((((........)).)))).((..(((.....(((((((.((((((....)))))).))).))))((((((........-)))))).)))..)).....------ ( -30.20, z-score = -1.06, R) >droSec1.super_2 7219455 97 + 7591821 UUCCC---CCCCACUGUGGAAAAUUGGGUCAAUGUGCUUUGAACUGAUUAGUGUCAGUCUAGUGGCAGUUGCCCGUUUCCU-GUCCACAACCUGAUCAAGG------ .....---......((((((....((((.((((.(((((((.((((((....)))))).))).))))))))))))......-.)))))).(((.....)))------ ( -28.90, z-score = -1.96, R) >droYak2.chr3L 7898370 93 + 24197627 -------GCCCCACCGUGGAAAAGUGGGUCAAUGCGCUGUGAACUGAUUAGUGUCAGUCUAGUGGCAGUGGCCCGUUUCCU-GGCCACAACCUGAUCAAGG------ -------......(((.(((((...((((((.(((((((.((.(((((....))))))))))).))).)))))).))))))-))......(((.....)))------ ( -32.80, z-score = -1.47, R) >droEre2.scaffold_4784 21714586 96 - 25762168 ----UUUUCCCCCCCGUGGAAAAGUGGGUCAAUGUGCUGUGAACUGAUUAGUGUCAGUCUAGUGGCAGUGGCCCGUUUCCU-GGCCACAACCUGAUCAAGG------ ----((((((.......))))))...(((((...(((..(..((((((....))))))...)..)))((((((........-))))))....)))))....------ ( -31.80, z-score = -1.70, R) >droAna3.scaffold_13337 14749332 86 + 23293914 ----UUGCUCCUCUGGUGGCUAAGUGGGUCAAUGUGCCCUGAACUGAUUAGUGUCAGUCUAGUGCCAGUGGCCAGUGGCCUGUUUCCUGG----------------- ----.....((...((.(((((....(((((.((.((.(((.((((((....)))))).))).)))).)))))..))))).....)).))----------------- ( -30.10, z-score = -1.47, R) >dp4.chrXR_group8 3384593 103 + 9212921 ----CCUGCCACCUUUCGGCCAAGUGGGUCAAUGUGCUGUGAACUGAUUUGUGUCAGUCUGGUGCUUGUGGCCCGCUUCCUUAUCCGCAACCUGAUCGAACCGAAGG ----........(((((((..((((((((((..(..(((.((.(((((....))))))))))..)...))))))))))....(((........)))....))))))) ( -34.10, z-score = -1.89, R) >droVir3.scaffold_13049 2454542 99 + 25233164 --UUCGCCCCAGCUGGCGUCAAAGUGGGUCAAUGUGCGUUGAACUGAUUAGCGCCUGG-UAUUCACCUUGGCUGGCCAAGGUGCGCGUAAUUAUGCCGCACA----- --......(((((((((((...(((.(((((((....)))).))).))).))))).((-(....)))..)))))).....(((((.(((....)))))))).----- ( -34.40, z-score = -0.17, R) >consensus ____CU_CCCCCCCUGUGGAAAAGUGGGUCAAUGUGCUUUGAACUGAUUAGUGUCAGUCUAGUGGCAGUGGCCCGUUUCCU_GGCCACAACCUGAUCAAGG______ ...............((((.....(((((((.((((((....((((((....))))))..))).))).))))))).........))))................... (-16.41 = -17.05 + 0.64)
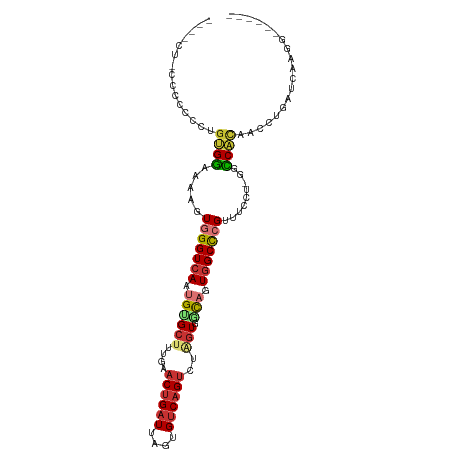


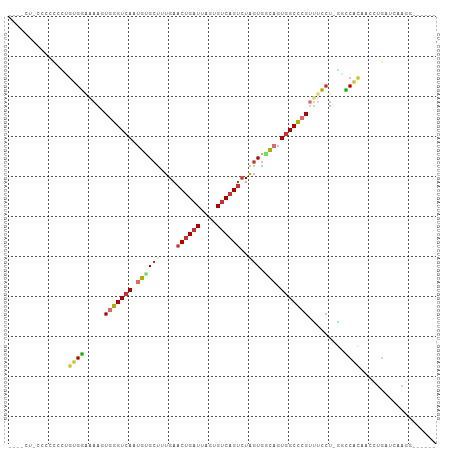
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:09:21 2011