
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 7,218,267 – 7,218,357 |
| Length | 90 |
| Max. P | 0.662004 |

| Location | 7,218,267 – 7,218,357 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 55.94 |
| Shannon entropy | 0.81080 |
| G+C content | 0.40001 |
| Mean single sequence MFE | -18.76 |
| Consensus MFE | -8.67 |
| Energy contribution | -7.46 |
| Covariance contribution | -1.20 |
| Combinations/Pair | 2.14 |
| Mean z-score | -0.37 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.662004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 7218267 90 - 24543557 -ACAUGGC-AGCUUUGUCAUUCG--GUCGUCUUCUUUGAAUGGCAAGGAUAUUUCAA-CGCAAGCUGCAUACAAUUUUGG--AUUUGAUUAAAUAGG---------------- -.....((-((((((((((((((--(.........))))))))))..((....))..-...)))))))...(((......--..)))..........---------------- ( -20.40, z-score = -1.18, R) >droPer1.super_1933 5451 106 - 6327 ACCCCAACGAACUAUAUUAUUUG-UCCCCUUGCUUUCAGAUGAUGAGGAUAUCUCGACAGCAAAGUGCAUUCGAACAUG----CAUACAUAAAUACGGAAUUG--AAGGCAGG ......(((((........))))-)...(((((((((((.((.((((.....)))).)).....((((((......)))----)))..............)))--)))))))) ( -21.80, z-score = -0.85, R) >dp4.chrXR_group6 3980947 112 - 13314419 ACCCCAACGAACUAUAUUAUUUG-UCCCCUUGCUUUCAGAGGAUGAGGAUAUCUCGACAGCAAAGUGCAUUCGAACAUGCAUACAUACAUAAAUACGGAAUUGCUAUAUGAGG .((.((.(((..(((....((..-(((.((.......)).)))..)).)))..)))..(((((.((((((......)))))).....(........)...)))))...)).)) ( -18.40, z-score = 0.43, R) >droAna3.scaffold_13337 1743627 87 - 23293914 -ACACGAC-AACUUUGUCAUCUGCCAUUUGCAUCUGAAAAUGGCAAAGACAUUCGAAGCGGCAACUCCAUGCA-UUUUGC--AUUAAAUGGG--------------------- -....(((-(....))))...((((((((........))))))))....((((..(.((((((......))).-...)))--.)..))))..--------------------- ( -19.80, z-score = -0.97, R) >droEre2.scaffold_4784 21637246 90 + 25762168 -ACAUGGC-AGCUUUGUCAUUCG--GUCGCCUUCUUUGAAUGGCAAGUGUAUUUCAA-CGCGAACUGUACACACUUUUGG--AUUUGAUUAAAUAGG---------------- -.....((-((..((((((((((--(.........)))))))))))((((......)-)))...))))............--...............---------------- ( -17.30, z-score = 0.27, R) >droYak2.chr3L 7818753 90 - 24197627 -ACAUGGC-AGCUUUGUCAUUCG--GUCGCCUUCUUUGAAUGGCAAGGAUAUUUCAA-CGCGAACUGUACACACUUUUGG--AUUUGAUGAAGUAGG---------------- -.......-..((((((((((((--(.........))))))))))))).(((((((.-..(((((((..........)))--.)))).)))))))..---------------- ( -19.50, z-score = -0.07, R) >droSec1.super_2 7144664 90 - 7591821 -ACAUGGC-AGCUUUGUCAUUCG--GUCGCCUUCUUUGAAUGGCAAGCAUAUUUCAG-CGCAAGCUGCAUACACUUUUGG--AUUUGAUUAAAUAGG---------------- -..(((..-.((((.((((((((--(.........)))))))))))))......(((-(....)))))))..........--...............---------------- ( -22.00, z-score = -0.72, R) >droSim1.chr3L 6726217 90 - 22553184 -ACAUGGC-AGUUUUGUCAUUCG--GUCGCCUUCUUUGAAUGGCAAGGAUAUUUCAA-CGCAAGCUGCAUACACUUUUGG--AUUUGAUUAAAUAGG---------------- -.....((-((((((((((((((--(.........))))))))))..((....))..-...)))))))............--...............---------------- ( -17.90, z-score = -0.08, R) >droGri2.scaffold_15110 12866362 86 - 24565398 -AUAUAAGAAAUAUUUCAAAGAUACUCCUGCAACUGUAUAUGGAAAAAGUACUCUG--GAAUUACCAUAUGGAAGCUU----CUUAAAAUGAG-------------------- -...((((((..((((((.((.(((((((((....)))...)))....))))).))--))))..((....))....))----)))).......-------------------- ( -11.70, z-score = -0.15, R) >consensus _ACAUGGC_AGCUUUGUCAUUCG__GUCGCCUUCUUUGAAUGGCAAGGAUAUUUCAA_CGCAAACUGCAUACAAUUUUGG__AUUUGAUUAAAUAGG________________ ...........((((((((((((.............))))))))))))...........((.....))............................................. ( -8.67 = -7.46 + -1.20)

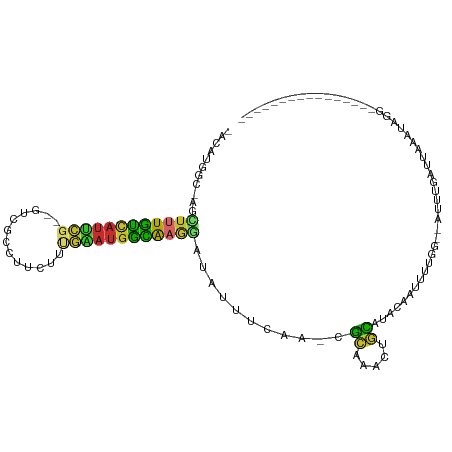

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:09:12 2011