
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 4,485,411 – 4,485,502 |
| Length | 91 |
| Max. P | 0.983410 |

| Location | 4,485,411 – 4,485,502 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 67.75 |
| Shannon entropy | 0.52767 |
| G+C content | 0.43651 |
| Mean single sequence MFE | -25.43 |
| Consensus MFE | -10.79 |
| Energy contribution | -13.97 |
| Covariance contribution | 3.19 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.983410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
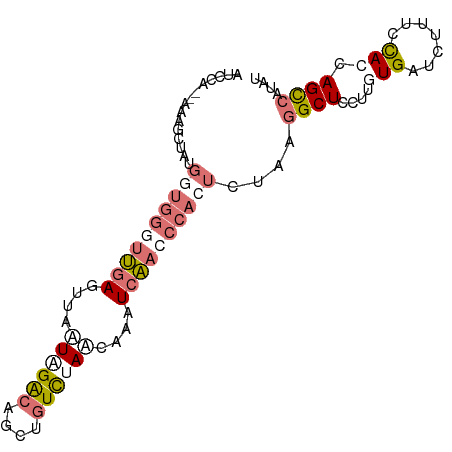
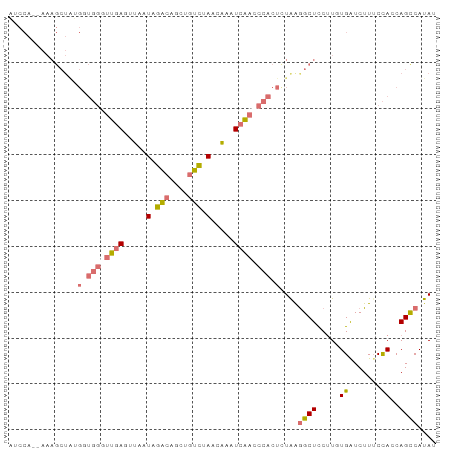
>dm3.chr2L 4485411 91 + 23011544 UACCA--AAAGCUGGGGUGGGUUGAGCUAUUGGACAGCUGUCUAACAAAUCAACCCACUCUAAGGCUUCUUUUGGUCUUUCCACCAGCCAUAU .....--...(((((((((((((((....((((((....))))))....))))))))))..((((((......))))))....)))))..... ( -36.20, z-score = -4.44, R) >droEre2.scaffold_4929 4575253 93 + 26641161 AUCCACCAAAGCUAUGGUGGAUUGAGUUAAUAGACAGCUGUCUAACAAAUCAACCCACUUUAAGGCUCCUUGUGAUCUUUCCACCAGUCAUAU .(((((((......)))))))((((.....(((((....))))).....))))..........((((....(((.......))).)))).... ( -21.70, z-score = -1.82, R) >droYak2.chr2L 4521577 91 + 22324452 AUCCA--AAAGCUAUGUUGGGUUGAGUUAAUAGACAUCUGUCCAACAAAUCAACCCACUCUGGAGCUCCUUUUGAUCUUUCCACCAGCCAUAU ...((--((((((..(.((((((((.......(((....))).......))))))))..)...))))...))))................... ( -20.04, z-score = -1.65, R) >droSec1.super_5 2584260 72 + 5866729 --------------------UCCCAAAAGAUAGG-GUGGGUUGAUCUAAUCGACUCACUCUAGGGCUCCAUGUGAUCUUACUACCAGCCAUAU --------------------..........((((-((((((((((...)))))))))))))).((((....(((....)))....)))).... ( -23.80, z-score = -2.49, R) >consensus AUCCA__AAAGCUAUGGUGGGUUGAGUUAAUAGACAGCUGUCUAACAAAUCAACCCACUCUAAGGCUCCUUGUGAUCUUUCCACCAGCCAUAU ...............((((((((((....((((((....))))))....))))))))))....((((....(((.......))).)))).... (-10.79 = -13.97 + 3.19)

| Location | 4,485,411 – 4,485,502 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 67.75 |
| Shannon entropy | 0.52767 |
| G+C content | 0.43651 |
| Mean single sequence MFE | -25.00 |
| Consensus MFE | -15.79 |
| Energy contribution | -18.60 |
| Covariance contribution | 2.81 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927761 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
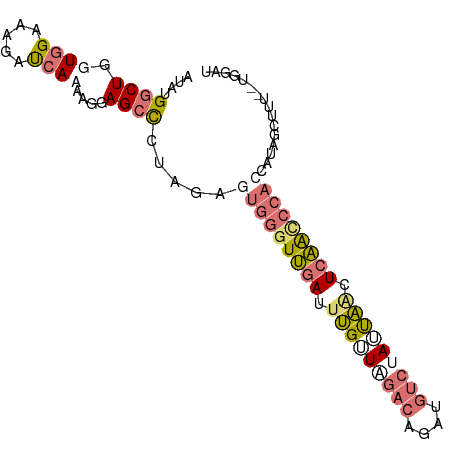
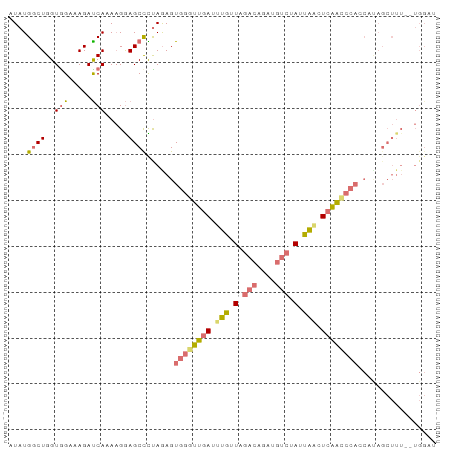
>dm3.chr2L 4485411 91 - 23011544 AUAUGGCUGGUGGAAAGACCAAAAGAAGCCUUAGAGUGGGUUGAUUUGUUAGACAGCUGUCCAAUAGCUCAACCCACCCCAGCUUU--UGGUA ....(((((((((.....)))..............(((((((((.(((((.(((....))).))))).))))))))).))))))..--..... ( -31.90, z-score = -2.71, R) >droEre2.scaffold_4929 4575253 93 - 26641161 AUAUGACUGGUGGAAAGAUCACAAGGAGCCUUAAAGUGGGUUGAUUUGUUAGACAGCUGUCUAUUAACUCAAUCCACCAUAGCUUUGGUGGAU ..................((((...((((......(((((((((.(((.(((((....))))).))).)))))))))....))))..)))).. ( -26.40, z-score = -1.54, R) >droYak2.chr2L 4521577 91 - 22324452 AUAUGGCUGGUGGAAAGAUCAAAAGGAGCUCCAGAGUGGGUUGAUUUGUUGGACAGAUGUCUAUUAACUCAACCCAACAUAGCUUU--UGGAU .................(((...(((((((......((((((((.(((.(((((....))))).))).))))))))....))))))--).))) ( -27.10, z-score = -1.89, R) >droSec1.super_5 2584260 72 - 5866729 AUAUGGCUGGUAGUAAGAUCACAUGGAGCCCUAGAGUGAGUCGAUUAGAUCAACCCAC-CCUAUCUUUUGGGA-------------------- ....((.((((.....((((((.(((....)))..)))).))......)))).))..(-((........))).-------------------- ( -14.60, z-score = 0.95, R) >consensus AUAUGGCUGGUGGAAAGAUCAAAAGGAGCCCUAGAGUGGGUUGAUUUGUUAGACAGAUGUCUAUUAACUCAACCCACCAUAGCUUU__UGGAU ....((((.((((.....))))....)))).....(((((((((.(((((((((....))))))))).)))))))))................ (-15.79 = -18.60 + 2.81)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:16:38 2011