
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,928,024 – 6,928,123 |
| Length | 99 |
| Max. P | 0.838595 |

| Location | 6,928,024 – 6,928,123 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 73.96 |
| Shannon entropy | 0.53301 |
| G+C content | 0.51040 |
| Mean single sequence MFE | -31.21 |
| Consensus MFE | -16.51 |
| Energy contribution | -18.08 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.838595 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3L 6928024 99 - 24543557 CAUUUUCCCGACAUUUCAGCU-UUUUCCCCCAAAAAACGAUGCGACGUCGCUGAUUGGAAGCUUCGGCAAACAGC--UCGUUUUUGGGGCGGAGCAUAAGCU .................((((-(.(((((((.((((((((.((......(((((..(....).))))).....))--)))))))).))).))))...))))) ( -31.10, z-score = -1.63, R) >droPer1.super_38 700385 91 - 801819 ----UUCCCGGCU---UCGC---UUUCCCCCAAAAA-CGAUGCGUCGUUGCUGAUUGGAAGCUUCGGCGCUUGCCUUUCGUUUUUGGGGCGGAGCAAAAGCU ----.....((((---(.((---((..(((((((((-(((.((((((..(((.......)))..)))))).......))))))))))))..))))..))))) ( -42.50, z-score = -3.79, R) >dp4.chrXR_group8 4223443 91 + 9212921 ----UUCCCGGCU---UCGC---UUUCCCCCAAAAA-CGAUGCGUCGUUGCUGAUUGGAAGCUUCGGCGCUUGCCUUUCGUUUUUGGGGCGGAGCAAAAGCU ----.....((((---(.((---((..(((((((((-(((.((((((..(((.......)))..)))))).......))))))))))))..))))..))))) ( -42.50, z-score = -3.79, R) >droAna3.scaffold_13337 3316782 96 + 23293914 -CACUUCCCGGCUCUGCCAC---UUUCCCCCAAAAAGCAAUGCGACGUCGCUGAUUGGAAGCUUCGGCACCCGGC--UCGGUUUUGGGGCGGAGCAAAAGCU -.........((((((((..---......(((((((((...(((....)))....(((..((....))..)))))--)...))))))))))))))....... ( -31.80, z-score = 0.21, R) >droYak2.chr3L 7510697 98 - 24197627 --CUUUCCCGACUUUUCUGCCUUUUUCCCCCAAAAAACGAUGCGACGUCGCUGAUUGGAAGCUUCGGCAAACAGC--UCGUUUUUGGGGCGGAGCAUAAACU --...........(((.(((....((((((((((((.(((.((......(((((..(....).))))).....))--)))))))))))).)))))).))).. ( -27.10, z-score = -0.66, R) >droSec1.super_2 6854798 99 - 7591821 CACUUUCCCGACUUUUCAGCU-UUUUCCCUCAAAAAACGAUGCGACGUCGCUGAUUGGAAGCUUCGGCAAACAGU--UCGUUUUUGGGGCGGAGCAUAAGCU .................((((-(.(((((((.((((((((.(((....)))..((((...((....))...))))--)))))))).))).))))...))))) ( -28.80, z-score = -1.03, R) >droSim1.chr3L 6401112 99 - 22553184 CACUUUCCCGACUUUUCAGCC-UUUUCCCCCAAAAAACGAUGCGACGUCGCUGAUUGGAAGCUUCGGCAAACAGC--UCGUUUUUGGGGCGGAGCAUAAGCU .................(((.-..(((((((.((((((((.((......(((((..(....).))))).....))--)))))))).))).)))).....))) ( -27.50, z-score = -0.29, R) >triCas2.ChLG8 9711038 91 - 15773733 -CACUUUUCGAAUUCUUUGUAGAUACGAUCUA-----CG-UGCCCAGAAGCUGAAAUGUAUUUUAUGCAAAUUGG--UUGUGAUC--GGCGAACGAUAUGCU -.((.(((((..((((.(((((((...)))))-----))-.....))))..))))).)).......(((.((((.--((((....--.)))).)))).))). ( -18.40, z-score = -0.11, R) >consensus __CUUUCCCGACUUUUCAGC__UUUUCCCCCAAAAAACGAUGCGACGUCGCUGAUUGGAAGCUUCGGCAAACAGC__UCGUUUUUGGGGCGGAGCAUAAGCU .......................(((((((((((((.(((.(((....))).........((....)).........)))))))))))).))))........ (-16.51 = -18.08 + 1.56)

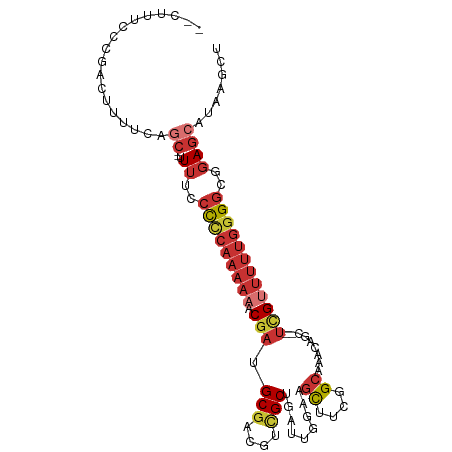

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:08:25 2011