| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,915,983 – 6,916,080 |
| Length | 97 |
| Max. P | 0.991915 |

| Location | 6,915,983 – 6,916,080 |
|---|---|
| Length | 97 |
| Sequences | 8 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 77.63 |
| Shannon entropy | 0.42151 |
| G+C content | 0.41478 |
| Mean single sequence MFE | -14.79 |
| Consensus MFE | -8.57 |
| Energy contribution | -8.97 |
| Covariance contribution | 0.41 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.51 |
| SVM RNA-class probability | 0.991915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
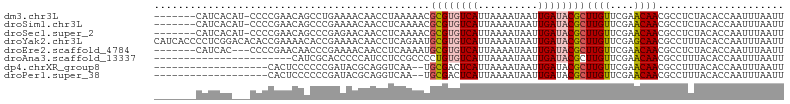
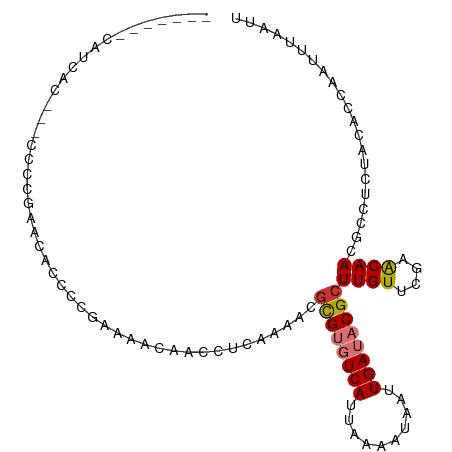
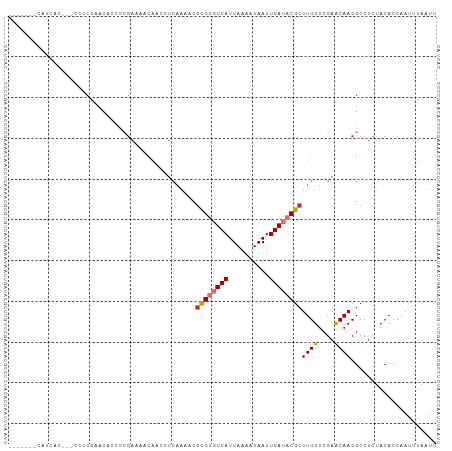
>dm3.chr3L 6915983 97 - 24543557 -------CAUCACAU-CCCCGAACAGCCUGAAAACAACCUAAAAACGCGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUCUACACCAAUUUAAUU -------........-...(((((((..((....))..........((((((((..........))))))))))))))).......................... ( -16.50, z-score = -3.57, R) >droSim1.chr3L 6389344 97 - 22553184 -------CAUCACAU-CCCCGAACAGCCCGAAAACAACCUCAAAACGCGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUCUACACCAAUUUAAUU -------........-...(((((((...((........)).....((((((((..........))))))))))))))).......................... ( -17.80, z-score = -4.06, R) >droSec1.super_2 6837991 97 - 7591821 -------CAUCACAU-CCCCGAACAGCCCGAGAACAACCUCAAAACGCGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUCUACACCAAUUUAAUU -------........-...(((((((...(((......))).....((((((((..........))))))))))))))).......................... ( -21.10, z-score = -4.94, R) >droYak2.chr3L 7497828 105 - 24197627 CAUCACCCCUCGGACACACCGAAAACACCGAAAACAACCUCAGAAUGCGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAGCAACGCCUUUACACCAAUUUAAUU ........((((((((...((.......))................((((((((..........)))))))).))))))))........................ ( -20.40, z-score = -3.18, R) >droEre2.scaffold_4784 21310846 95 + 25762168 -------CAUCAC---CCCCGAACAACCCGAAAACAACCUCAAAAUGCGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUCUACACCAAUUUAAUU -------......---...(((((((...((........)).....((((((((..........))))))))))))))).......................... ( -17.80, z-score = -4.80, R) >droAna3.scaffold_13337 3302367 82 + 23293914 -----------------------CAUCGCACCCCCAUCCUCCGCCCCUGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUUUACACCAAUUUAAUU -----------------------.........................((((((..........))))))((((((....)))).)).................. ( -8.10, z-score = -1.29, R) >dp4.chrXR_group8 4211054 84 + 9212921 -------------------CACUCCCCCCGAUACGCAGGUCAA--UGCGACUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUUUACACCAAUUUAAUU -------------------..........((..((((......--))))..)).................((((((....)))).)).................. ( -8.30, z-score = 0.16, R) >droPer1.super_38 687934 84 - 801819 -------------------CACUCCCCCCGAUACGCAGGUCAA--UGCGACUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUUUACACCAAUUUAAUU -------------------..........((..((((......--))))..)).................((((((....)))).)).................. ( -8.30, z-score = 0.16, R) >consensus _______CAUCAC___CCCCGAACACCCCGAAAACAACCUCAAAACGCGUGUCAUUAAAAUAAUUGAUACGCUUGUUCGAACAACGCCUCUACACCAAUUUAAUU ..............................................((((((((..........))))))))((((....))))..................... ( -8.57 = -8.97 + 0.41)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:08:23 2011