
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,908,875 – 6,908,965 |
| Length | 90 |
| Max. P | 0.829884 |

| Location | 6,908,875 – 6,908,965 |
|---|---|
| Length | 90 |
| Sequences | 7 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 73.97 |
| Shannon entropy | 0.48895 |
| G+C content | 0.47050 |
| Mean single sequence MFE | -28.12 |
| Consensus MFE | -15.92 |
| Energy contribution | -15.74 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.829884 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 6908875 90 - 24543557 AACCACAGCAG--GCUUAUGCCUGCGGUCGCUUCGAUUGCAUCAGAUGA-GGUGCGA--------------AUCC-CAAUCGGAAGCUAAAUUAGUGCUAAAAAAAAA ...(((.((((--((....))))))....((((((((((((((......-)))))(.--------------....-))))).))))).......)))........... ( -28.20, z-score = -2.32, R) >droSim1.chr3L 6381779 91 - 22553184 AACCGCAGCAGACGCUUAUGCCUGCGGUCGCUGCGAUUGCAUCAGAUGA-GGUGCGA--------------AUCC-CAAUCGGAAGCUAAAUCAGUGCUAAAAAAAA- ...((((((.(((((........)).))))))))).(((((((......-)))))))--------------.(((-.....)))(((.........)))........- ( -27.40, z-score = -1.10, R) >droSec1.super_2 6830833 92 - 7591821 AACCGCAGCAGACGCUUAUGCCUGCGGUCGCUGCGAUUGCAUCAGAUGA-GGUGCGA--------------AUCC-CAAUCGGAAGCUAAAUCAGUGCUAAAAAAAAA ...((((((.(((((........)).))))))))).(((((((......-)))))))--------------.(((-.....)))(((.........)))......... ( -27.40, z-score = -1.15, R) >droYak2.chr3L 7489182 105 - 24197627 AACCGCAGCAGACGCUUAUGCCUGCGGUCGCCUCGAUUGCAACAGAUGA-GGUGCGAAAUCGCAAGCGAGCAUCC-CAAUCGGAAGCUAAAUCAGUGCUAGAAAAAA- .(((((((((........)).))))))).((((((.(((...))).)))-)))(((....))).(((((((.(((-.....))).))).......))))........- ( -31.31, z-score = -0.76, R) >droEre2.scaffold_4784 21303447 106 + 25762168 AACCGCAGCAGACGCUUAUGCCUGCGGUCGCGUCGAUUGCAUCAGAUGA-GGUGCGAAAUCGCAAUCGAGCAUCC-CAAUCGGAAGCUAAAACAGUGCUAAAAAAUAA .(((((((((........)).))))))).(((((((((((.((....))-(((.....)))))))))))((.(((-.....))).))........))).......... ( -31.70, z-score = -1.18, R) >droAna3.scaffold_13337 3294856 83 + 23293914 AACCGCAGCAGACGCUUAUGCCUGCGGUUGUUCGGAUUGCAUCAGAUGA-GGUGCAG--------------AUCCAGAAUCAGAUUCUCAAUCGGUUC---------- ((((((((((........)).))))))))....((((((((((......-)))))).--------------)))).(((((.(((.....))))))))---------- ( -29.50, z-score = -2.30, R) >droWil1.scaffold_180916 2482110 99 + 2700594 AACCACAACAAACGCUUAUGAUUGCGGUUGUGUUUAUUGCAUCAGAUGAUGAUGAAGAUGAGAAUGUA---AAUUGAAAUCAGAAUCUGAAUCCGAAUCAAA------ ...((((((...(((........))))))))).........((((((..((((...(((.........---.)))...))))..))))))............------ ( -21.30, z-score = -1.24, R) >consensus AACCGCAGCAGACGCUUAUGCCUGCGGUCGCUUCGAUUGCAUCAGAUGA_GGUGCGA______________AUCC_CAAUCGGAAGCUAAAUCAGUGCUAAAAAAAA_ .(((((((((........)).)))))))........(((((((.......)))))))................................................... (-15.92 = -15.74 + -0.18)


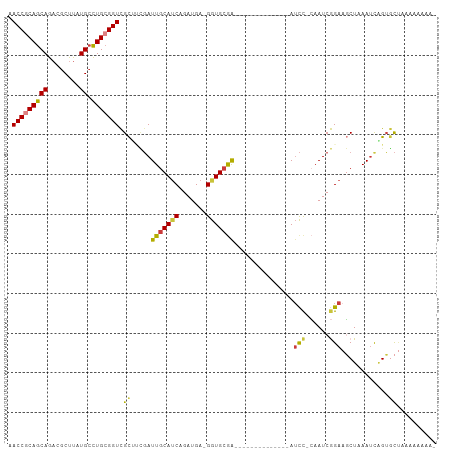
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:08:23 2011