
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,866,402 – 6,866,524 |
| Length | 122 |
| Max. P | 0.755121 |

| Location | 6,866,402 – 6,866,493 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 77.60 |
| Shannon entropy | 0.42948 |
| G+C content | 0.43039 |
| Mean single sequence MFE | -19.90 |
| Consensus MFE | -11.76 |
| Energy contribution | -11.57 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.545507 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
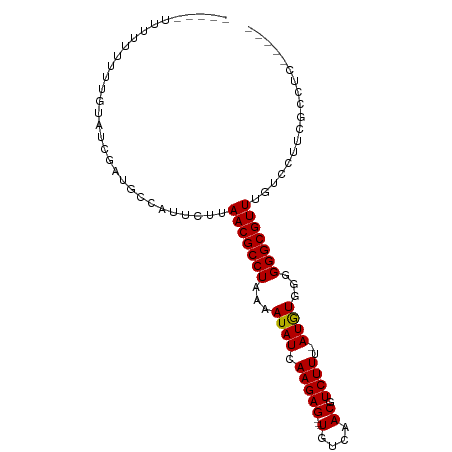
>dm3.chr3L 6866402 91 + 24543557 UUUUCUUUUUUUUUGUAUCGAUGCCAUUCUUAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUGUGCGGGGCGUUUGUCCUUCGCUUC----- ...................((.((.......(((((((...((((.(((((---(....)).)))).-))))...)))))))........)).))----- ( -18.16, z-score = -1.42, R) >droSim1.chr3L 6345368 86 + 22553184 -----UCCUUUUUUGUAUCGAUGCCAUUCUUAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUGUGCGGGGCGUUUGUCCUUCGCCUC----- -----..............((.((.......(((((((...((((.(((((---(....)).)))).-))))...)))))))........)).))----- ( -18.16, z-score = -1.29, R) >droSec1.super_2 6796045 86 + 7591821 -----UCCUUUUUUGUAUCGAUGCCAUUCCUAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUGUGCGGGGCGUUUGUCCUUCGCCUC----- -----..............((.((.......(((((((...((((.(((((---(....)).)))).-))))...)))))))........)).))----- ( -18.16, z-score = -1.27, R) >droYak2.chr3L 7451572 88 + 24197627 ---UUUUUUUUUUUGUAUCGCUGCCAUUCUUAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUGUGGGGGGCGUUAGUCCUUCGCCUC----- ---...............((.((.(((((((...............)))))---)).)).)).....-..(((((((((....)))))))))...----- ( -19.26, z-score = -1.32, R) >droEre2.scaffold_4784 21267441 83 - 25762168 --------UUUUUUGUAUCGAUGCCAUUCUUAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUAUGGGGGGCGUUUGUCCUUCGCUUC----- --------...........((.((.......(((((((...((((.(((((---(....)).)))).-))))...)))))))........)).))----- ( -18.16, z-score = -1.27, R) >droPer1.super_38 642855 84 + 801819 --------GUGCUUUUUUUUAUACCAUUCUUAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUGUGGGGGGCGUUUGUCCUUAAAGGCC---- --------..((((((....(((........(((((((...((((.(((((---(....)).)))).-))))...))))))))))....)))))).---- ( -19.00, z-score = -1.07, R) >droWil1.scaffold_180916 2426398 90 - 2700594 ------UGUUGUUCUGUGUCGCCCCAUAUAAAACGCCUAAAAUAUCAAGAG---UGUCAACGUCUUU-AUGUUGGGGGCGUUUGUCCAGCCAGCCAGCUC ------.((((..(((.((((((((.......((((((.........)).)---)))((((((....-))))))))))))........))))).)))).. ( -23.50, z-score = -1.64, R) >droVir3.scaffold_13049 7216773 91 + 25233164 ---UUUUUUUAGUGGCUGUGAUAC-GUUCUUAACGCCUAAAAUAUCAAGAGCAAUGUCAACGUCUUUGAUAUGGGGGGCGUUUACCCC--AAGGCCC--- ---..........((((.((....-((....(((((((...((((((((..(.........)..))))))))...))))))).))..)--).)))).--- ( -24.80, z-score = -1.19, R) >consensus _____UUUUUUUUUGUAUCGAUGCCAUUCUUAACGCCUAAAAUAUCAAGAG___UGUCAACGUCUUU_AUGUGGGGGGCGUUUGUCCUUCGCCUC_____ ...............................(((((((...((((...(((...((....)).)))..))))...))))))).................. (-11.76 = -11.57 + -0.19)


| Location | 6,866,431 – 6,866,524 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 69.69 |
| Shannon entropy | 0.62579 |
| G+C content | 0.46261 |
| Mean single sequence MFE | -24.42 |
| Consensus MFE | -12.02 |
| Energy contribution | -12.05 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.13 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.755121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
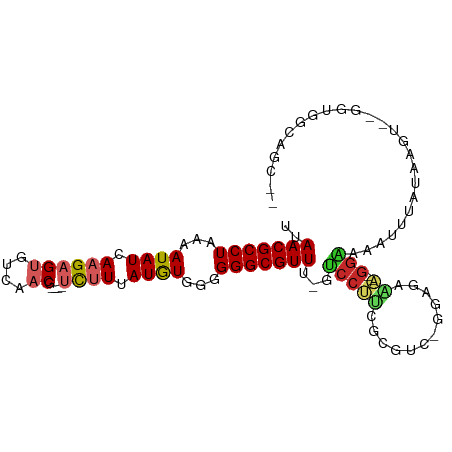
>dm3.chr3L 6866431 93 + 24543557 UUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUGUGCGGGGCGUUU-GUCCUUCGCUUC-GGAGAAA--GGAAAAUUUACAAGU--GGUGGCAGC-- ..(((((((...((((.((((((....)).----)))).))))...)))))))(-(((..((((((.-(...(((--......))).)))))--)).))))..-- ( -23.90, z-score = -0.80, R) >droSim1.chr3L 6345392 93 + 22553184 UUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUGUGCGGGGCGUUU-GUCCUUCGCCUC-GGAGAAA--GGAAAAUUUAUAAGU--GGUGGCAGC-- ..(((((((...((((.((((((....)).----)))).))))...))))))).-...(((((((.(-....(((--......)))....).--))))).)).-- ( -22.40, z-score = -0.31, R) >droSec1.super_2 6796069 93 + 7591821 CUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUGUGCGGGGCGUUU-GUCCUUCGCCUC-GGAGAAA--GGAAAAUUUAUAAGU--GGUGGCAGC-- ..(((((((...((((.((((((....)).----)))).))))...))))))).-...(((((((.(-....(((--......)))....).--))))).)).-- ( -22.40, z-score = -0.29, R) >droYak2.chr3L 7451598 92 + 24197627 UUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUGUGGGGGGCGUUA-GUCCUUCGCCUC-GGAGAAA--GGAAA-UUUAUAAGU--GGUGGCAGC-- (((((((((...((((.((((((....)).----)))).))))...))))))))-)..(((((((.(-....(((--.....-)))....).--))))).)).-- ( -24.40, z-score = -1.11, R) >droEre2.scaffold_4784 21267462 92 - 25762168 UUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUAUGGGGGGCGUUU-GUCCUUCGCUUC-GGAGAAA--GGAAA-UUUAUAAGU--GGUGGCAGC-- ..(((((((...((((.((((((....)).----)))).))))...))))))).-.((((((.(...-.).).))--)))..-.........--.........-- ( -23.50, z-score = -1.23, R) >droAna3.scaffold_13337 3248946 99 - 23293914 UUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCCUUAUGUGGGGGGCGUUUUGUCCUUCGAGGCAGGAGAAAUGGGCAAACCUUCGAGACAGGAGGGAGA-- .....(((((................((((----(((((.....)))))))))((.((((.......)))).)).)))))...(((((......)))))....-- ( -28.70, z-score = -0.65, R) >droPer1.super_38 642876 93 + 801819 UUAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUGUGGGGGGCGUUU-GUCCUUAAAGGCCAGAGGGA--GGGAAAUCGAAGAAAAGGGCGAC----- ...((((((...((((.((((((....)).----)))).))))...))))))((-((((((.....((....)).--((....))......)))))))).----- ( -23.80, z-score = -0.95, R) >droWil1.scaffold_180916 2426421 98 - 2700594 AAAACGCCUAAAAUAUCAAGAGUGUCAACG----UCUUUAUGUUGGGGGCGUUU-GUCCAGCCAGCCAGCUCACU--AAAAGAUAAGAAAAAAAAGGGAGAGUCC .((((((((..(((((.((((((....)).----)))).)))))..))))))))-.((((((......)))..((--........)).........)))...... ( -22.30, z-score = -0.99, R) >droVir3.scaffold_13049 7216798 80 + 25233164 UUAACGCCUAAAAUAUCAAGAGCAAUGUCAACGUCUUUGAUAUGGGGGGCGUUU-ACCCC-AAGGCCCGAUGCGA--GGGGGAU--------------------- ..(((((((...((((((((..(.........)..))))))))...))))))).-.((((-...((.....))..--.))))..--------------------- ( -27.90, z-score = -1.97, R) >droMoj3.scaffold_6680 5483105 91 + 24764193 UUAACGCCUAAAAUAUCAAGAGCAAUGUC------UCUGAUAUGGGGGGCGUUU-ACCCC-AAGGGCUGAUUGGA--GGGAGAUACAACCACACACACACA---- ..(((((((...((((((.(((......)------))))))))...))))))).-.((((-((.......)))).--))......................---- ( -24.90, z-score = -1.59, R) >consensus UUAACGCCUAAAAUAUCAAGAGUGUCAACG____UCUUUAUGUGGGGGGCGUUU_GUCCUUCGCGUC_GGAGAAA__GGAAAAUUUAUAAGU__GGUGGCAGC__ ..(((((((...((((.((((.............)))).))))...))))))).................................................... (-12.02 = -12.05 + 0.03)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:08:16 2011