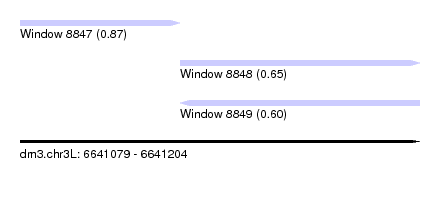
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,641,079 – 6,641,204 |
| Length | 125 |
| Max. P | 0.874325 |

| Location | 6,641,079 – 6,641,129 |
|---|---|
| Length | 50 |
| Sequences | 4 |
| Columns | 50 |
| Reading direction | forward |
| Mean pairwise identity | 86.67 |
| Shannon entropy | 0.20980 |
| G+C content | 0.39500 |
| Mean single sequence MFE | -10.98 |
| Consensus MFE | -7.84 |
| Energy contribution | -8.65 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.874325 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 6641079 50 + 24543557 ACGUUUCAAUUAAAAUGAAGACAAAGGUGGUCUGGAAAACAGGACGGCAA ..((((((.......)).))))....((.((((........)))).)).. ( -8.30, z-score = -0.89, R) >droSim1.chr3L 6113257 50 + 22553184 AGGUUUCAAUUAAAAUUGAAACAAAGGUGCUCUGGAAAACAGGACGGCAA ..((((((((....))))))))...(.((..(((.....)))..)).).. ( -13.70, z-score = -3.48, R) >droSec1.super_2 6577907 50 + 7591821 AGGUUUCAAUUAAAAUUGAAACAAAGGUGCUCUGGAAAACAGGACGGCAA ..((((((((....))))))))...(.((..(((.....)))..)).).. ( -13.70, z-score = -3.48, R) >droYak2.chr3L 7214930 50 + 24197627 ACGUUUCAAUUAAAAUGGAGACAAAGCUAGUCGGGAACACAGGAGAGCAU ..(((((..........)))))...(((..((.(.....).))..))).. ( -8.20, z-score = -1.71, R) >consensus ACGUUUCAAUUAAAAUGGAAACAAAGGUGCUCUGGAAAACAGGACGGCAA ..((((((((....))))))))...(.((..(((.....)))..)).).. ( -7.84 = -8.65 + 0.81)

| Location | 6,641,129 – 6,641,204 |
|---|---|
| Length | 75 |
| Sequences | 10 |
| Columns | 87 |
| Reading direction | forward |
| Mean pairwise identity | 72.04 |
| Shannon entropy | 0.53540 |
| G+C content | 0.43741 |
| Mean single sequence MFE | -19.10 |
| Consensus MFE | -10.77 |
| Energy contribution | -9.02 |
| Covariance contribution | -1.75 |
| Combinations/Pair | 1.73 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.653330 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
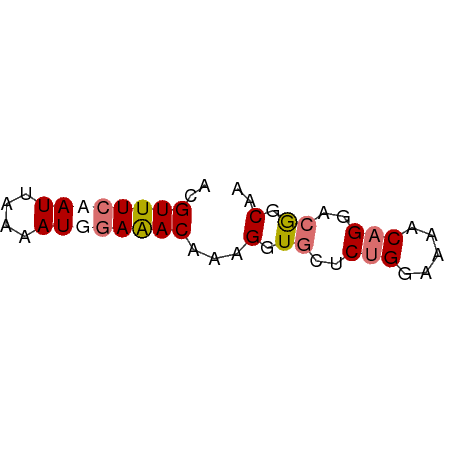
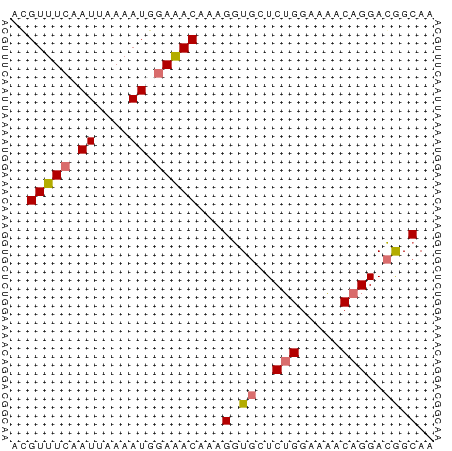
>dm3.chr3L 6641129 75 + 24543557 CUGGAAUGGCCGUUGAAACUUUUUUCCAAGUGCCAUUUGGCAUACAAAGAG---------AGAUGAUA---AGAGUGAAACACUCAA ..((.....)).......(((((((....(((((....)))))...)))))---------))......---.(((((...))))).. ( -16.40, z-score = -0.99, R) >droSim1.chr3L 6113307 75 + 22553184 AUGGAAUGGCCGUUGAAACUUUUUUCCAAGUGCCAUUUGGCAUACAAAGAG---------AGAUGAUA---AGAGUGAAACACUCAA ((((.....)))).....(((((((....(((((....)))))...)))))---------))......---.(((((...))))).. ( -17.10, z-score = -1.44, R) >droSec1.super_2 6577957 75 + 7591821 CUGGAAUGGCCGUUGAAACUUUUUUCCAAGUGCCAUUUGGCAUACAAAGAG---------AGAUGAUA---AGAGUGAAACACUCAA ..((.....)).......(((((((....(((((....)))))...)))))---------))......---.(((((...))))).. ( -16.40, z-score = -0.99, R) >droYak2.chr3L 7214980 75 + 24197627 CUGGGAUGGCCGUUGAAACUUUUUUUCAAGUGCCAUUUGGCAUACAAAGAC---------AGAUGAUA---AGAGUGAAACACUCAA ((((((((((..((((((.....))))))..)))))))(.....).....)---------))......---.(((((...))))).. ( -18.10, z-score = -1.42, R) >droEre2.scaffold_4784 9316399 77 + 25762168 CUGGGAUGGCCAUUGAAACUUUUUUCCAAGUGCCAUUCGGCAUACAUAGAGGG-------AGAUGAUA---AGAGUGAAACACUCAA .(((.....)))......(((((((....(((((....))))).....)))))-------))......---.(((((...))))).. ( -16.10, z-score = -0.14, R) >droAna3.scaffold_13337 3000042 80 - 23293914 AGGAAGUGGUUGCGGCAACUUUUUUCCAAGUACCAUUUGGCAUACAAUAAAGG-------AGGGAUUCUUAAGAGCAAACCACUUAA ...(((((((((((....).(((((((..(((((....))..)))......))-------))))).........)).)))))))).. ( -18.40, z-score = -1.01, R) >dp4.chrXR_group8 3911681 87 - 9212921 CGGAAGUGGUUGCGGCAACUUUUUGCCGAGUACCAUUUGGCAUAAAAAAGGCAGAAGUGUGGAUAAGAGAGAGAGCGAGCCACUCAA ..((.(((((((((((((....))))).....(((((((.(........).)))....))))............)).)))))))).. ( -22.90, z-score = -1.06, R) >droPer1.super_38 392030 87 + 801819 CGGAAGUGGUUGCGGCAACUUUUUGCCGAGUACCAUUUGGCAUAAAAAAGGCAGAAGUGUGGAUAAGAGAGAGAGCGAGCCACUCAA ..((.(((((((((((((....))))).....(((((((.(........).)))....))))............)).)))))))).. ( -22.90, z-score = -1.06, R) >droMoj3.scaffold_6680 14950763 76 - 24764193 CGGAAGUGGUUGCAGCAACUUUU-GCCGAGUGCCAUUUGGCACACAAAAGAU--------GGAUGACGU--CGACUGAGACAGACAA (((....(((((...)))))...-.))).(((((....))))).((...(((--------(.....)))--)...)).......... ( -20.40, z-score = -0.83, R) >droGri2.scaffold_15110 16256803 75 - 24565398 CGGAAGUGGUUGCAGCAACUUUU-GCCGAGUGCCAUUUGGCACACAAAAGAU--------GGAUGACGC--UGAAUAAGGCAGUCA- (((....(((((...)))))...-.))).(((((....))))).........--------...((((((--(......))).))))- ( -22.30, z-score = -1.07, R) >consensus CGGAAAUGGCCGCUGAAACUUUUUUCCAAGUGCCAUUUGGCAUACAAAAAG_________AGAUGAUA___AGAGUGAAACACUCAA ...(((((((..(.((((....)))).)...)))))))..................................(((.......))).. (-10.77 = -9.02 + -1.75)



| Location | 6,641,129 – 6,641,204 |
|---|---|
| Length | 75 |
| Sequences | 10 |
| Columns | 87 |
| Reading direction | reverse |
| Mean pairwise identity | 72.04 |
| Shannon entropy | 0.53540 |
| G+C content | 0.43741 |
| Mean single sequence MFE | -16.29 |
| Consensus MFE | -7.65 |
| Energy contribution | -7.21 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.602983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
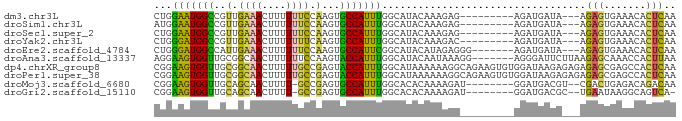
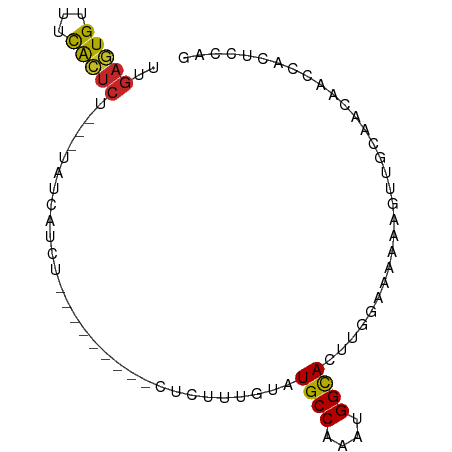
>dm3.chr3L 6641129 75 - 24543557 UUGAGUGUUUCACUCU---UAUCAUCU---------CUCUUUGUAUGCCAAAUGGCACUUGGAAAAAAGUUUCAACGGCCAUUCCAG ..(((((...))))).---........---------..............((((((..((((((.....))))))..)))))).... ( -14.70, z-score = -1.18, R) >droSim1.chr3L 6113307 75 - 22553184 UUGAGUGUUUCACUCU---UAUCAUCU---------CUCUUUGUAUGCCAAAUGGCACUUGGAAAAAAGUUUCAACGGCCAUUCCAU ..(((((...))))).---........---------..............((((((..((((((.....))))))..)))))).... ( -14.70, z-score = -1.26, R) >droSec1.super_2 6577957 75 - 7591821 UUGAGUGUUUCACUCU---UAUCAUCU---------CUCUUUGUAUGCCAAAUGGCACUUGGAAAAAAGUUUCAACGGCCAUUCCAG ..(((((...))))).---........---------..............((((((..((((((.....))))))..)))))).... ( -14.70, z-score = -1.18, R) >droYak2.chr3L 7214980 75 - 24197627 UUGAGUGUUUCACUCU---UAUCAUCU---------GUCUUUGUAUGCCAAAUGGCACUUGAAAAAAAGUUUCAACGGCCAUCCCAG ..(((((...))))).---........---------...............(((((..((((((.....))))))..)))))..... ( -14.30, z-score = -1.17, R) >droEre2.scaffold_4784 9316399 77 - 25762168 UUGAGUGUUUCACUCU---UAUCAUCU-------CCCUCUAUGUAUGCCGAAUGGCACUUGGAAAAAAGUUUCAAUGGCCAUCCCAG ..(((((...))))).---........-------...............(.(((((..((((((.....))))))..))))).)... ( -13.90, z-score = -0.43, R) >droAna3.scaffold_13337 3000042 80 + 23293914 UUAAGUGGUUUGCUCUUAAGAAUCCCU-------CCUUUAUUGUAUGCCAAAUGGUACUUGGAAAAAAGUUGCCGCAACCACUUCCU ..((((((((.((..............-------......(((.....)))..(((((((......))).))))))))))))))... ( -17.00, z-score = -1.55, R) >dp4.chrXR_group8 3911681 87 + 9212921 UUGAGUGGCUCGCUCUCUCUCUUAUCCACACUUCUGCCUUUUUUAUGCCAAAUGGUACUCGGCAAAAAGUUGCCGCAACCACUUCCG ..((((((...((................................((((....))))...(((((....)))))))..))))))... ( -18.50, z-score = -1.92, R) >droPer1.super_38 392030 87 - 801819 UUGAGUGGCUCGCUCUCUCUCUUAUCCACACUUCUGCCUUUUUUAUGCCAAAUGGUACUCGGCAAAAAGUUGCCGCAACCACUUCCG ..((((((...((................................((((....))))...(((((....)))))))..))))))... ( -18.50, z-score = -1.92, R) >droMoj3.scaffold_6680 14950763 76 + 24764193 UUGUCUGUCUCAGUCG--ACGUCAUCC--------AUCUUUUGUGUGCCAAAUGGCACUCGGC-AAAAGUUGCUGCAACCACUUCCG ((((..(((......)--))......(--------(.((((((((((((....)))))...))-))))).))..))))......... ( -17.70, z-score = -1.17, R) >droGri2.scaffold_15110 16256803 75 + 24565398 -UGACUGCCUUAUUCA--GCGUCAUCC--------AUCUUUUGUGUGCCAAAUGGCACUCGGC-AAAAGUUGCUGCAACCACUUCCG -((((.((........--))))))..(--------(.((((((((((((....)))))...))-))))).))............... ( -18.90, z-score = -1.55, R) >consensus UUGAGUGUUUCACUCU___UAUCAUCU_________CUCUUUGUAUGCCAAAUGGCACUUGGAAAAAAGUUGCAACAACCACUCCAG ..(((((...)))))..............................((((....)))).............................. ( -7.65 = -7.21 + -0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:07:34 2011