| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,587,042 – 6,587,103 |
| Length | 61 |
| Max. P | 0.974144 |

| Location | 6,587,042 – 6,587,103 |
|---|---|
| Length | 61 |
| Sequences | 12 |
| Columns | 64 |
| Reading direction | reverse |
| Mean pairwise identity | 75.72 |
| Shannon entropy | 0.52593 |
| G+C content | 0.36617 |
| Mean single sequence MFE | -12.15 |
| Consensus MFE | -9.75 |
| Energy contribution | -9.11 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.974144 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3L 6587042 61 - 24543557 ACUCUCGUGAUAGU-CCUUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGCUGUACACAUU-- ......(((((((.-.(((...(((......)))(((......))))))..)))).)))...-- ( -13.20, z-score = -0.84, R) >droSim1.chr3L 6058726 63 - 22553184 ACUCUCGUGAUAGU-CCUUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGCUGUACACAUUAA ......(((((((.-.(((...(((......)))(((......))))))..)))).)))..... ( -13.20, z-score = -0.76, R) >droSec1.super_2 6523618 63 - 7591821 ACUCUCGUGAUAGU-CCUUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGCUGUACACAUUAA ......(((((((.-.(((...(((......)))(((......))))))..)))).)))..... ( -13.20, z-score = -0.76, R) >droYak2.chr3L 7158921 61 - 24197627 ACUCUCGUGAUAGU-CCUUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGCUGUACACAUU-- ......(((((((.-.(((...(((......)))(((......))))))..)))).)))...-- ( -13.20, z-score = -0.84, R) >droEre2.scaffold_4784 9261210 61 - 25762168 ACUCUCGUGAUAGU-CCUUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGCUGUACUCAUU-- .(((((((......-...(((((((......))).)))).....)))))))...........-- ( -11.84, z-score = -0.43, R) >droAna3.scaffold_13337 2942980 59 + 23293914 AAUCUCGUGAUG---UCCUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAUCUGUACACGUU-- .(((((((.((.---...(((((((......))).))))...)))))))))...........-- ( -10.70, z-score = -0.35, R) >dp4.chrXR_group8 3855302 57 + 9212921 ACUCCCGCGAUUG--UCCUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGAAAU-CAUA---- .(((.((((.(((--(...((((((......))).))))))).)))).)))....-....---- ( -12.10, z-score = -1.69, R) >droPer1.super_38 335446 57 - 801819 ACUCCCGCGAUUA--UCCUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGAAAU-CAUA---- .(((.((((.(((--(...((((((......))).))))))).)))).)))....-....---- ( -12.10, z-score = -2.00, R) >droWil1.scaffold_180916 1912530 54 + 2700594 --UCUCGCUAUUG--UCUUUUGUACGUUUUAGUAGCAAAUAAAUGCGAGAUACGCUUU------ --((((((.(((.--...(((((((......)).)))))..)))))))))........------ ( -13.10, z-score = -2.02, R) >droMoj3.scaffold_6680 14886422 50 + 24764193 CAUCUCCUCGUUGU--ACUUUGUACGUUUUAGUAGCAAAU-AAAACGAGUUUU----------- ......((((((..--..(((((((......)).))))).-..))))))....----------- ( -10.50, z-score = -2.68, R) >droGri2.scaffold_15110 16194949 62 + 24565398 AAUUUCGUGAUUGUCUUUUUUGUACGUUUUAGUAGCAAAUAAAAACGAGAUUCGUCACUUUU-- ......(((((.(((((((((((..(((.....)))..))))))..)))))..)))))....-- ( -10.80, z-score = -1.06, R) >droVir3.scaffold_13049 10946179 62 - 25233164 AAUCUCGUGAUUGU-CUUUUUGUACGUUUUAGUAGCAAAUAAAAACGAGAUUCUUAACAUUCU- ((((((((..((((-....((((((......)).))))))))..))))))))...........- ( -11.80, z-score = -1.97, R) >consensus ACUCUCGUGAUUGU_CCUUUUGUGCGUUUUAGUAGCAAAUAAAUGCGAGAGCUGUACACAUU__ .(((((((..........(((((((......))).)))).....)))))))............. ( -9.75 = -9.11 + -0.64)

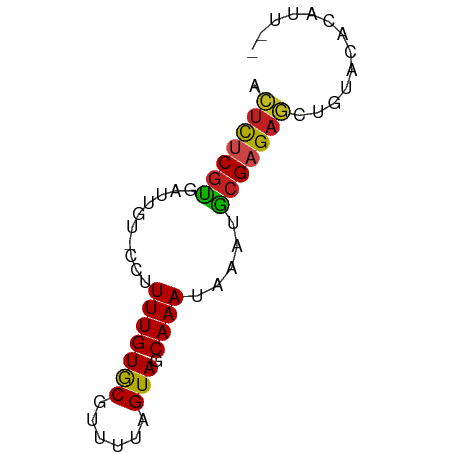
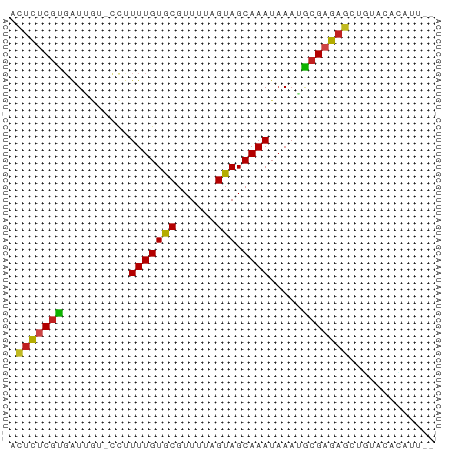
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:07:29 2011