
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,497,524 – 6,497,619 |
| Length | 95 |
| Max. P | 0.992419 |

| Location | 6,497,524 – 6,497,619 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 75.62 |
| Shannon entropy | 0.45315 |
| G+C content | 0.44520 |
| Mean single sequence MFE | -25.00 |
| Consensus MFE | -17.89 |
| Energy contribution | -18.20 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.54 |
| SVM RNA-class probability | 0.992419 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
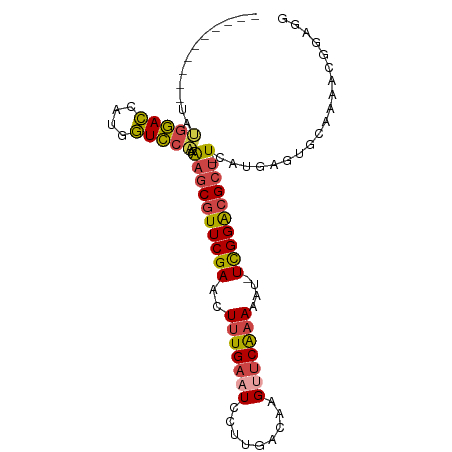
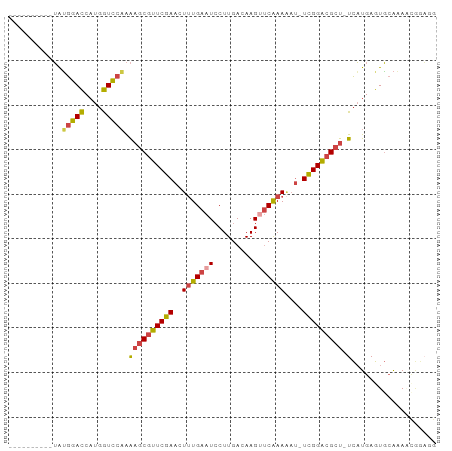
>dm3.chr3L 6497524 95 + 24543557 UAUGGACCAGUGCGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGCUCAAAAAAAUUGGACGCU-UCAUGAGUGCAAAACGGAGG ......((..(((((((....))))..((((((((((..(((((.............)))))....)))))))))-).......)))....))... ( -24.12, z-score = -0.89, R) >droSim1.chr3L 5968707 83 + 22553184 ----------UAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAAAAU-UCGGGCGCU-UCAUGAGUGCAAA-CGGAGG ----------..(((((....))))).(((((((((((.(((((((.........)))))))..)-)))))))))-)............-...... ( -25.40, z-score = -2.28, R) >droSec1.super_2 6434719 84 + 7591821 ----------UAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAAAAU-UCGGACGCU-UCAUGAGUGCAAAACGGAGG ----------..(((((....))))).(((((((((((.(((((((.........)))))))..)-)))))))))-)................... ( -25.90, z-score = -2.83, R) >droEre2.scaffold_4784 9167599 91 + 25762168 ---CGACCAAUAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAAAAU-UCGGACGCU-UCAUGAGUGCAAAAUGGAGG ---...(((...(((((....))))).(((((((((((.(((((((.........)))))))..)-)))))))))-).............)))... ( -28.90, z-score = -3.25, R) >droYak2.chr3L 7066171 84 + 24197627 ---CAACCUAUAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAAUAU-UCGGACGCU-UCAUGAGUGCAAA------- ---.........(((((....))))).(((((((((((.(((((((.........)))))))..)-)))))))))-)............------- ( -25.70, z-score = -3.69, R) >droWil1.scaffold_180916 1767677 86 - 2700594 ---------CAGGACAUCAUCAUGAUGGUUCAUUCGAACUGUGAAUGCUUGACAAGCCCGAAGUU-UCGGACGCUAUUUUGAAUUGGCAGCAGUAG ---------((((.(((..((((...(((((....))))))))))))))))....((((((....-))))..((((........)))).))..... ( -20.00, z-score = -0.04, R) >consensus __________UAUGGACCAUGGUCCAAAAGCGUUCGAACUUUGAAUCCUUGACAAGUUCAAAAAU_UCGGACGCU_UCAUGAGUGCAAAACGGAGG ............(((((....)))))..(((((((((..(((((((.........)))))))....)))))))))..................... (-17.89 = -18.20 + 0.31)

| Location | 6,497,524 – 6,497,619 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 75.62 |
| Shannon entropy | 0.45315 |
| G+C content | 0.44520 |
| Mean single sequence MFE | -24.40 |
| Consensus MFE | -16.97 |
| Energy contribution | -17.37 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.54 |
| SVM RNA-class probability | 0.992367 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
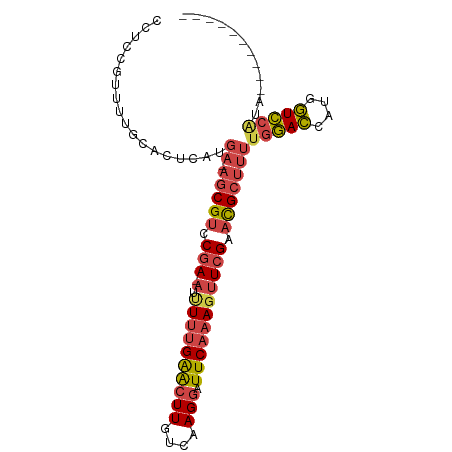
>dm3.chr3L 6497524 95 - 24543557 CCUCCGUUUUGCACUCAUGA-AGCGUCCAAUUUUUUUGAGCUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCGCACUGGUCCAUA .....(((((........))-)))((((((....(((((((((............))))))))).....)))))).((((.((.....)).)))). ( -24.10, z-score = -1.03, R) >droSim1.chr3L 5968707 83 - 22553184 CCUCCG-UUUGCACUCAUGA-AGCGCCCGA-AUUUUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUA---------- ......-...........((-((((..(((-(((((.((((((....))).)))))))))))..))))))(((((....)))))..---------- ( -24.60, z-score = -2.57, R) >droSec1.super_2 6434719 84 - 7591821 CCUCCGUUUUGCACUCAUGA-AGCGUCCGA-AUUUUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUA---------- ..................((-(((((.(((-(((((.((((((....))).))))))))))).)))))))(((((....)))))..---------- ( -26.60, z-score = -3.26, R) >droEre2.scaffold_4784 9167599 91 - 25762168 CCUCCAUUUUGCACUCAUGA-AGCGUCCGA-AUUUUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUAUUGGUCG--- ...(((............((-(((((.(((-(((((.((((((....))).))))))))))).)))))))(((((....)))))...)))...--- ( -28.40, z-score = -3.17, R) >droYak2.chr3L 7066171 84 - 24197627 -------UUUGCACUCAUGA-AGCGUCCGA-AUAUUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUAUAGGUUG--- -------...........((-(((((.(((-((.(((((((((....))).))))))))))).)))))))(((((....))))).........--- ( -26.60, z-score = -3.26, R) >droWil1.scaffold_180916 1767677 86 + 2700594 CUACUGCUGCCAAUUCAAAAUAGCGUCCGA-AACUUCGGGCUUGUCAAGCAUUCACAGUUCGAAUGAACCAUCAUGAUGAUGUCCUG--------- .....((((...........))))((((((-....))))))........(((((.......)))))...((((.....)))).....--------- ( -16.10, z-score = 0.08, R) >consensus CCUCCGUUUUGCACUCAUGA_AGCGUCCGA_AUUUUUGAACUUGUCAAGGAUUCAAAGUUCGAACGCUUUUGGACCAUGGUCCAUA__________ .....................(((((.(((...((((((((((....))).))))))).))).)))))..(((((....)))))............ (-16.97 = -17.37 + 0.40)
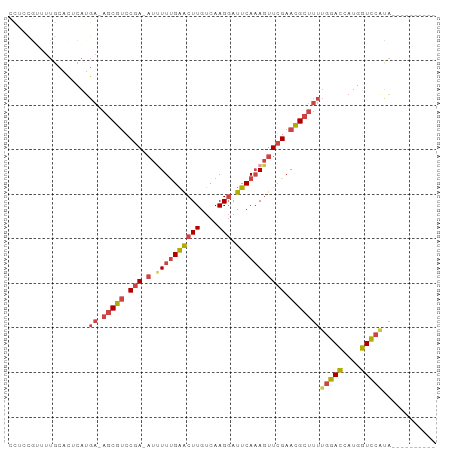
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:07:11 2011