
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 6,103,442 – 6,103,538 |
| Length | 96 |
| Max. P | 0.768250 |

| Location | 6,103,442 – 6,103,538 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 76.71 |
| Shannon entropy | 0.44586 |
| G+C content | 0.55311 |
| Mean single sequence MFE | -26.32 |
| Consensus MFE | -22.24 |
| Energy contribution | -21.79 |
| Covariance contribution | -0.45 |
| Combinations/Pair | 1.20 |
| Mean z-score | -0.76 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.768250 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
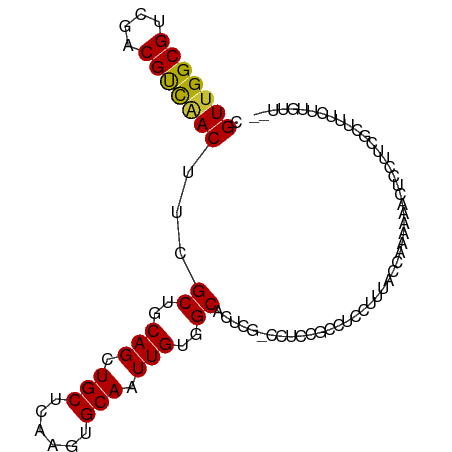
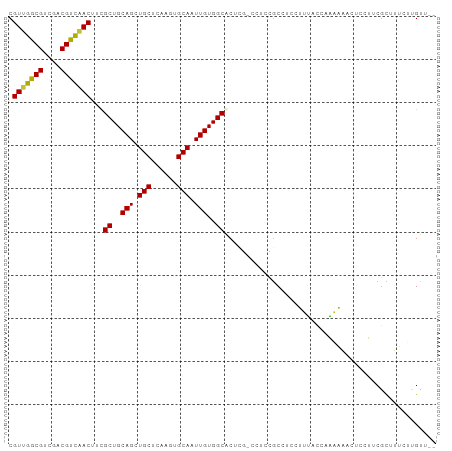
>dm3.chr3L 6103442 96 - 24543557 CGUUGGCGUCGACGUCAACUUCGCUGCAGCUGCUCAAGUGCAAUUGUGGCACUCG-CCUCCGCCUCCUUUACCAAAAAUCUCCUUCGCUUUCUUGUU-- .(((((((....)))))))...((..(((.(((......))).)))..)).....-.........................................-- ( -22.30, z-score = -1.09, R) >droSim1.chr3L 5606851 96 - 22553184 CGUUGGCGUCGACGUCAACUUCGCUGCAGCUGCUCAAGUGCAAUUGUGGCACUCG-CCUCCGCCUCCUUUACCAAAAAUCUCCUUCGUUUUCUUGUU-- .(((((((....)))))))...((..(((.(((......))).)))..)).....-.........................................-- ( -22.30, z-score = -1.30, R) >droSec1.super_2 6051646 97 - 7591821 CGUUGGCGUCGACGUCAACUUCGCUGCAGCUGCUCAAGUGCAAUUGUGGCACUCGCCCUCCGCCUCCUUUACCCAAAAUCUCCUUCGUUUUCUUGUU-- .(((((((....)))))))...((..(((.(((......))).)))..))...............................................-- ( -22.30, z-score = -1.45, R) >droYak2.chr3L 6687117 96 - 24197627 CGUUGGCGUCGACGUCAACUUUGCUGCAGCUGCUCAAGUGCAAUUGUGGCACUCG-CCUCUGCCUCCUUUUCCAAAAAACUCCUGCGCUUUCUUGUU-- .(((((((....)))))))...((.((((.(((......))).....((((....-....))))..................)))))).........-- ( -23.40, z-score = -0.70, R) >droEre2.scaffold_4784 8802251 96 - 25762168 CGUUGGCGUCGACGUCAACUUUGCUGCAGCUGCUCAAGUGCAAUUGUGGCACUCG-CCUCCGCCUCCUUUUCCAAAAAACUCCUUCGCUUUCUUGUU-- .(((((((....)))))))..(((..(((.(((......))).)))..)))....-.........................................-- ( -22.60, z-score = -1.02, R) >droAna3.scaffold_13337 2459963 98 + 23293914 CGUUGGCGUCGACGUUCACAUCGCUGCAGCUGCUCAAGUGCAAUUGUGGCGCUUA-CCCCUACCCUGGCUACUCGCCGACCAGCUCCUUUGCGCGUCCU .(((((((.............(((..(((.(((......))).)))..)))....-..((......)).....)))))))..((......))....... ( -27.50, z-score = 0.36, R) >dp4.chrXR_group6 3745994 93 + 13314419 CGUUGGCGUCUACGCUGACAGCGCUGCAGCUGCUCAAGUGCAAUUGUGGCGC----UGCCUGCCUCAUGCCUUUGUAA--CAGCGCGCCUUGUCGUCAU ((..((((....(((((.((((((..(((.(((......))).)))..))))----))...((.....))........--)))))))))....)).... ( -35.10, z-score = -0.43, R) >droPer1.super_35 155796 93 - 973955 CGUUGGCGUCUACGCUGACAGCGCUGCAGCUGCUCAAGUGCAAUUGUGGCGC----UGCCUGCCUCAUGCCUUUGUAA--CAGCGCGCCUUGUCGUCAU ((..((((....(((((.((((((..(((.(((......))).)))..))))----))...((.....))........--)))))))))....)).... ( -35.10, z-score = -0.43, R) >consensus CGUUGGCGUCGACGUCAACUUCGCUGCAGCUGCUCAAGUGCAAUUGUGGCACUCG_CCUCCGCCUCCUUUACCAAAAAACUCCUUCGCUUUCUUGUU__ .(((((((....)))))))...((..(((.(((......))).)))..))................................................. (-22.24 = -21.79 + -0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:06:34 2011