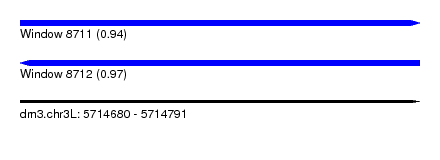

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,714,680 – 5,714,791 |
| Length | 111 |
| Max. P | 0.974139 |

| Location | 5,714,680 – 5,714,791 |
|---|---|
| Length | 111 |
| Sequences | 7 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 52.09 |
| Shannon entropy | 0.92645 |
| G+C content | 0.32034 |
| Mean single sequence MFE | -24.85 |
| Consensus MFE | 0.00 |
| Energy contribution | 0.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 0.00 |
| Mean z-score | -3.40 |
| Structure conservation index | -0.00 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.938655 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5714680 111 + 24543557 CAAAUAGGAUUAAAAAAGACUAAGUUUAAAUUGAAAGUAGACAGGAGCUGUGUGCAUUUUCUGUGCGCCCUGUCUAGUUUUUACUUAAAAGAGAAAGACUCUUUCAAAGUA--- ....(((..((....))..)))........(((((((((((((((....(((..((.....))..)))))))))))((((((.(((....))))))))).)))))))....--- ( -29.10, z-score = -2.57, R) >droSim1.chr3L 5226640 98 + 22553184 CAAAUAGGAUUAAAAAAGAUUAAGUUUAAAUUGAAAGUAGACAGAAGCUGUGUGC-------------CCUGUCUAGUUUUUACUUAAAAGAGAAAAACUCUUUCAAAGUA--- .((((..((((......))))..))))...((((((((((((((..((.....))-------------.)))))))((((((.(((....))))))))).)))))))....--- ( -21.10, z-score = -2.43, R) >droSec1.super_2 5650527 98 + 7591821 CAAAUAGGAUUAAAAAAGAUUAAGUUUAAAUUGAAAGUAGACAGGAGCUGUGUGC-------------CCUGUCUAGUUUUUACUUAAAAGAGAAAAACUCUUUCAAAGUA--- .((((..((((......))))..))))...(((((((((((((((.((.....))-------------))))))))((((((.(((....))))))))).)))))))....--- ( -26.20, z-score = -3.87, R) >droEre2.scaffold_4784 8405357 111 + 25762168 CAAAUAGGAUUAAAAAAGAUUAACUUUAAAUUGAAAGUGGACCAGAGUGGUGUGCAUUUUAUGUAUGCCCUGCUUAGAGUUCACUUAAAAAAGAAAAGCCCUUUCAAAGUA--- ......................(((((.......(((((((((.(((((((((((((...)))))))))..)))).).))))))))......((((.....))))))))).--- ( -24.70, z-score = -2.21, R) >droAna3.scaffold_13337 10466195 99 - 23293914 ----CAACGUCAAAGUGCAUCAGUCUUCAAUUAAAUACAAAUU-----UAUUGAAGCAUUAAGUACACCCUUCCAACAUCCUGUUGGAAAGA---AACUUUUCACAAAGUA--- ----.....((...((((...((((((((((.((((....)))-----)))))))).)))..))))....(((((((.....))))))).))---.(((((....))))).--- ( -19.70, z-score = -3.19, R) >dp4.chrXR_group6 5402252 107 - 13314419 ---CUAAUUUCGGGAUAAAAUAUGCCUUAUUUUGGCAUAAUCUAUA----CCAAACUUAGGCAUAUAUGUUCCUCGAAUACAACUUACAACAGACAACUGCUCAGAAAUCUGUU ---.....(((((((....((((((((...(((((.(((...))).----)))))...))))))))....))).))))..........((((((...((....))...)))))) ( -26.70, z-score = -4.79, R) >droPer1.super_27 548905 111 + 1201981 ---CUAAUUUCGGGAUAAAAUAUGCCUUAUUUUGGCAUAAUCUACAUACACCAAAUUUAGGCAUAUAUGUUCCUCGAAUACAACUUACAACAGACAACUGCUCAGAAAUCUGUU ---.....(((((((....((((((((...(((((...............)))))...))))))))....))).))))..........((((((...((....))...)))))) ( -26.46, z-score = -4.75, R) >consensus CAAAUAGGAUUAAAAAAGAUUAAGUUUAAAUUGAAAGUAGACAAGAGCUGUGUGC_UUUU__GUAUACCCUGCCUAGAUUUUACUUAAAAGAGAAAAACUCUUUCAAAGUA___ .................................................................................................................. ( 0.00 = 0.00 + 0.00)
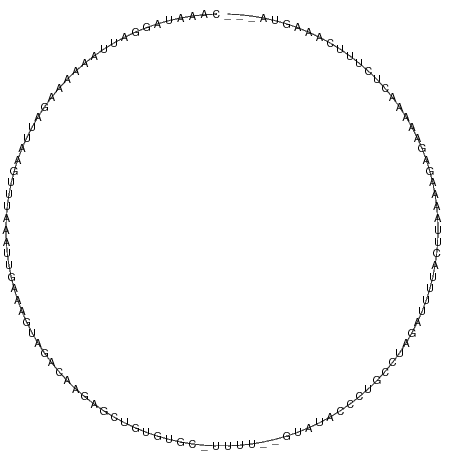


| Location | 5,714,680 – 5,714,791 |
|---|---|
| Length | 111 |
| Sequences | 7 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 52.09 |
| Shannon entropy | 0.92645 |
| G+C content | 0.32034 |
| Mean single sequence MFE | -26.34 |
| Consensus MFE | 0.00 |
| Energy contribution | 0.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 0.00 |
| Mean z-score | -3.66 |
| Structure conservation index | -0.00 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.974139 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5714680 111 - 24543557 ---UACUUUGAAAGAGUCUUUCUCUUUUAAGUAAAAACUAGACAGGGCGCACAGAAAAUGCACACAGCUCCUGUCUACUUUCAAUUUAAACUUAGUCUUUUUUAAUCCUAUUUG ---....(((((((((.((......(((((((..(((.(((((((((((((.......))).....)).)))))))).)))..)))))))...)).)))))))))......... ( -25.40, z-score = -3.08, R) >droSim1.chr3L 5226640 98 - 22553184 ---UACUUUGAAAGAGUUUUUCUCUUUUAAGUAAAAACUAGACAGG-------------GCACACAGCUUCUGUCUACUUUCAAUUUAAACUUAAUCUUUUUUAAUCCUAUUUG ---(((((.(((((((.....)))))))))))).....((((((((-------------((.....)).))))))))..................................... ( -21.00, z-score = -3.48, R) >droSec1.super_2 5650527 98 - 7591821 ---UACUUUGAAAGAGUUUUUCUCUUUUAAGUAAAAACUAGACAGG-------------GCACACAGCUCCUGUCUACUUUCAAUUUAAACUUAAUCUUUUUUAAUCCUAUUUG ---(((((.(((((((.....)))))))))))).....((((((((-------------((.....)).))))))))..................................... ( -24.00, z-score = -4.61, R) >droEre2.scaffold_4784 8405357 111 - 25762168 ---UACUUUGAAAGGGCUUUUCUUUUUUAAGUGAACUCUAAGCAGGGCAUACAUAAAAUGCACACCACUCUGGUCCACUUUCAAUUUAAAGUUAAUCUUUUUUAAUCCUAUUUG ---....(((((((.((((....(((......)))....)))).((((((.......))))..(((.....))))).))))))).((((((........))))))......... ( -16.40, z-score = -0.29, R) >droAna3.scaffold_13337 10466195 99 + 23293914 ---UACUUUGUGAAAAGUU---UCUUUCCAACAGGAUGUUGGAAGGGUGUACUUAAUGCUUCAAUA-----AAUUUGUAUUUAAUUGAAGACUGAUGCACUUUGACGUUG---- ---..(..(((.(((....---.(((((((((.....)))))))))(((((.(((...((((((((-----(((....)))).)))))))..))))))))))).)))..)---- ( -25.90, z-score = -2.91, R) >dp4.chrXR_group6 5402252 107 + 13314419 AACAGAUUUCUGAGCAGUUGUCUGUUGUAAGUUGUAUUCGAGGAACAUAUAUGCCUAAGUUUGG----UAUAGAUUAUGCCAAAAUAAGGCAUAUUUUAUCCCGAAAUUAG--- (((((((..((....))..)))))))..........((((.(((....((((((((...(((((----((((...)))))))))...))))))))....))))))).....--- ( -36.20, z-score = -5.94, R) >droPer1.super_27 548905 111 - 1201981 AACAGAUUUCUGAGCAGUUGUCUGUUGUAAGUUGUAUUCGAGGAACAUAUAUGCCUAAAUUUGGUGUAUGUAGAUUAUGCCAAAAUAAGGCAUAUUUUAUCCCGAAAUUAG--- (((((((..((....))..)))))))..........((((.(((....((((((((.(.(((((((((.......))))))))).).))))))))....))))))).....--- ( -35.50, z-score = -5.31, R) >consensus ___UACUUUGAAAGAGGUUUUCUCUUUUAAGUAAAAUCUAGGCAGGGUAUAC__AAAA_GCACACAGCUCCAGUCUACUUUCAAUUUAAACUUAAUCUUUUUUAAUCCUAUUUG .................................................................................................................. ( 0.00 = 0.00 + 0.00)
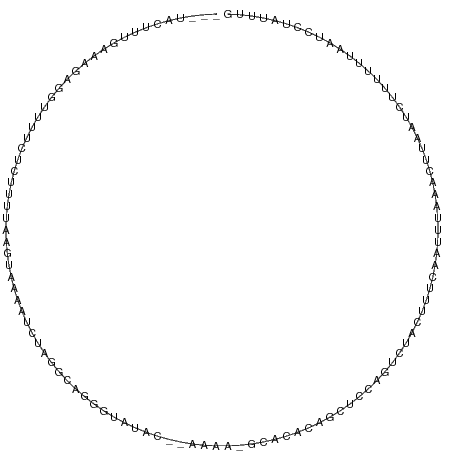


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:05:43 2011