
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,705,204 – 5,705,308 |
| Length | 104 |
| Max. P | 0.796354 |

| Location | 5,705,204 – 5,705,308 |
|---|---|
| Length | 104 |
| Sequences | 7 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 76.13 |
| Shannon entropy | 0.43992 |
| G+C content | 0.47799 |
| Mean single sequence MFE | -25.84 |
| Consensus MFE | -15.20 |
| Energy contribution | -16.34 |
| Covariance contribution | 1.14 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.796354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5705204 104 + 24543557 CGGUAACAGGCCAGUUCAGGCCAGGCUCACCGCU-CAACUGCAGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA--------- .(((....((((......))))..((.....(((-(....).))).....((((((....)))))))))))....(((((((.(((....))).)))))))....--------- ( -29.30, z-score = -1.59, R) >droSim1.chr3L 5217188 104 + 22553184 CGGGAAGAGGCCAGUUCAGGCCAGGCUCACCGCU-CAACUGCAGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA--------- .((..((.((((......))))...))..))(((-(....).))).....((((((....)))))).........(((((((.(((....))).)))))))....--------- ( -31.00, z-score = -1.67, R) >droSec1.super_2 5641134 96 + 7591821 --------GGCCAGUUCAGGCCAGGCUCACCGCU-CAACUGCAGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA--------- --------((((......))))..((.....(((-(....).))).....((((((....)))))))).......(((((((.(((....))).)))))))....--------- ( -28.30, z-score = -1.95, R) >droYak2.chr3L 6271976 85 + 24197627 -----------------CACC---GCUCACCGCUUCAACUGCAGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA--------- -----------------....---((.....(((.(....).))).....((((((....)))))))).......(((((((.(((....))).)))))))....--------- ( -21.10, z-score = -1.97, R) >droEre2.scaffold_4784 8396133 101 + 25762168 CGGGAACAGGCCAGGCUCACC---GCUCACCGCU-CAACUGCAGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA--------- .(((....((...(((.....---)))..))(((-(....).))).))).((((((....)))))).........(((((((.(((....))).)))))))....--------- ( -24.90, z-score = -0.38, R) >droAna3.scaffold_13337 10456197 90 - 23293914 -----------CCCCCUUAGCCAGGCAU-CCGCC-CACCU--UGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA--------- -----------........(((.(((..-..)))-.....--.........(((((....)))))))).......(((((((.(((....))).)))))))....--------- ( -22.40, z-score = -1.90, R) >droWil1.scaffold_180698 3964296 103 + 11422946 ----------CCAGAUCCCCUGCCGUUUGCCUCUCUCUCUCUUG-GCUCACACGUUAUAUAAUGCGCCAUCAUUUUGUGGUCAGCAUUUAUGCACCCCAAAAUGAAUGUGGACA ----------.(((.....)))..(((..(............((-((.((............)).))))((((((((.(((..(((....)))))).))))))))..)..))). ( -23.90, z-score = -1.47, R) >consensus ________GGCCAGUUCAAGCCAGGCUCACCGCU_CAACUGCAGCACUCACACGUUAUAUAAUGUGGCAUCAUUUUGUGGUCAGCAUUUAUGCAGACCAUAUACA_________ ........................((.....(((........))).....((((((....)))))))).......(((((((.(((....))).)))))))............. (-15.20 = -16.34 + 1.14)
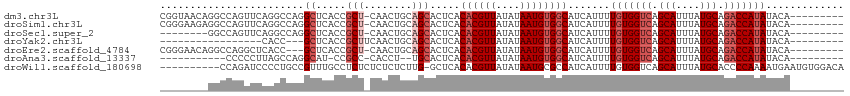
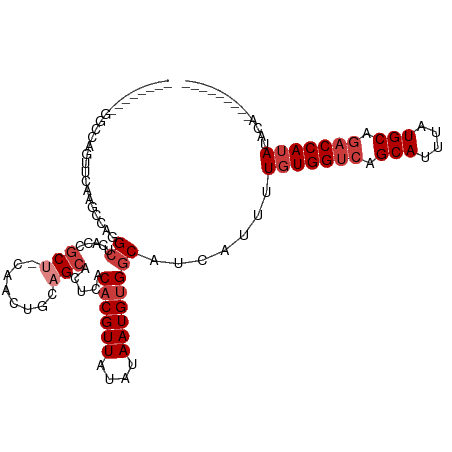
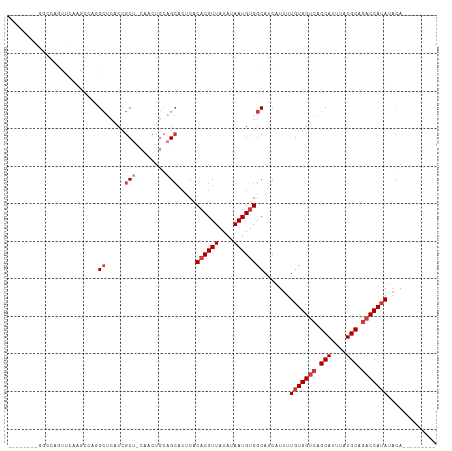
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:05:40 2011