
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 4,382,691 – 4,382,821 |
| Length | 130 |
| Max. P | 0.590418 |

| Location | 4,382,691 – 4,382,788 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.51 |
| Shannon entropy | 0.45801 |
| G+C content | 0.43441 |
| Mean single sequence MFE | -29.25 |
| Consensus MFE | -13.07 |
| Energy contribution | -15.70 |
| Covariance contribution | 2.63 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.526463 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
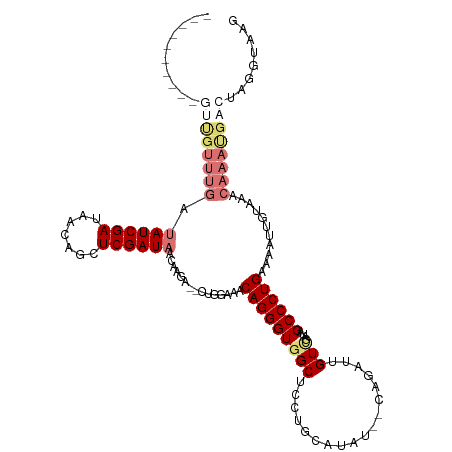
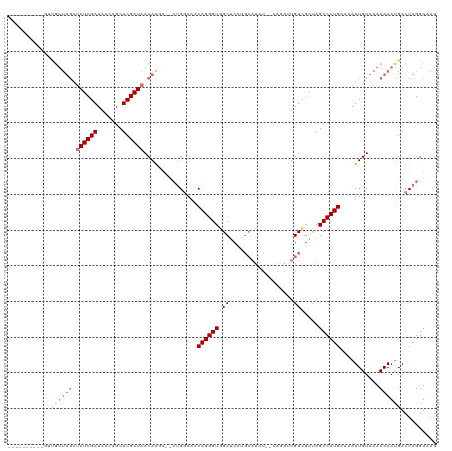
>dm3.chr2L 4382691 97 - 23011544 ----------AGCGUUUGAUAUCGAUAAGCGCUCGAUAACAAUACGCUGAAAACAGGGUUGCCCG-------------UUGUUGAUAGCCCUGAAAAUUGUAAACAAACGACUAGGUAAG ----------((((((((.((((((.......)))))).))).))))).....((((((((..((-------------....)).))))))))....((((......))))......... ( -24.50, z-score = -1.90, R) >droSim1.chr2L 4305709 116 - 22036055 AGCGUAUCUGGUUGUUUGGUAUCGAUAACAGCUCGAUAACAAGG--CUGGAAACAGGGUGGCUCCUGCAUAU--CAGAUUGUCGAUAGCCCUGAGCAUUGUAAACAAAUGACUAGGUAAG ....((((((((..((((.((((((.......))))))((((.(--(((....)(((((.((....)).(((--(........))))))))).))).))))...))))..)))))))).. ( -38.20, z-score = -2.71, R) >droSec1.super_5 2480957 101 - 5866729 -----------------AGUAUCGAUAACAGCUCGAUAACAAGAGUCUGGAAACAGGGUGGCUCCGGCAUAU--CAGAUUGUCGAUAGCCCUGAACAUUGUAAACAAAUGACUAGGUAAG -----------------..((((((((((.((((........))))(((....)))((((((....)).)))--).).)))))))))..((((..((((.......))))..)))).... ( -26.00, z-score = -1.55, R) >droEre2.scaffold_4929 4464950 116 - 26641161 GGGGUAUCUUGUUGUUUGAGAUCGAUAACAGCUCGAUAACAGAA----GGAGACAGGGUGGCUCGUGCAUAUAGAAAACUGUUCACAGCCCUGAAAAUUGUAAACAAAUGACUAGGUGAG ....(((((.((..((((..(((((.......))))).((((..----(....)..((.((((.(((...((((....)))).))))))))).....))))...))))..)).))))).. ( -28.30, z-score = -0.77, R) >consensus __________GUUGUUUGAUAUCGAUAACAGCUCGAUAACAAGA__CUGGAAACAGGGUGGCUCCUGCAUAU__CAGAUUGUCGAUAGCCCUGAAAAUUGUAAACAAAUGACUAGGUAAG ..........((((((((.((((((.......))))))...............(((((((((..................)))....))))))...........))))))))........ (-13.07 = -15.70 + 2.63)

| Location | 4,382,731 – 4,382,821 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 65.84 |
| Shannon entropy | 0.45875 |
| G+C content | 0.40620 |
| Mean single sequence MFE | -24.43 |
| Consensus MFE | -11.94 |
| Energy contribution | -11.83 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.590418 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
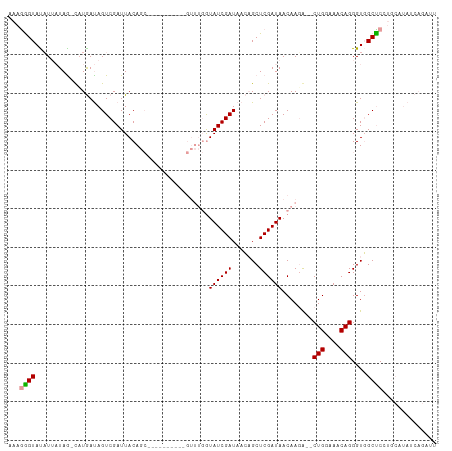
>dm3.chr2L 4382731 90 - 23011544 AAAGGGUAUAUUUUAGUUAUAACAGUCGAUUACAGC----------GUUUGAUAUCGAUAAGCGCUCGAUAACAAUACGCUGAAAACAGGGUUGCCCGUU----------- ...(((((.(((((.(((....(....)....((((----------(((((.((((((.......)))))).))).))))))..)))))))))))))...----------- ( -23.70, z-score = -2.20, R) >droSim1.chr2L 4305749 108 - 22036055 AAAGGGUAUUUUAUAA-CCUGAUAGUCGAUUACAGCGUAUCUGGUUGUUUGGUAUCGAUAACAGCUCGAUAACAAGG--CUGGAAACAGGGUGGCUCCUGCAUAUCAGAUU ....((((((..((((-((.((((((........)).)))).))))))..))))))((((.((((.(........))--)))....(((((....)))))..))))..... ( -28.20, z-score = -1.17, R) >droSec1.super_5 2480997 92 - 5866729 -AAAAGUACAUGAUAG-CAUGAUAGUCGAUUACA-----------------GUAUCGAUAACAGCUCGAUAACAAGAGUCUGGAAACAGGGUGGCUCCGGCAUAUCAGAUU -...............-..(((((((((......-----------------.((((((.......))))))....(((((((....)))....)))))))).))))).... ( -21.40, z-score = -1.46, R) >consensus AAAGGGUAUAUUAUAG_CAUGAUAGUCGAUUACAGC__________GUUUGGUAUCGAUAACAGCUCGAUAACAAGA__CUGGAAACAGGGUGGCUCCUGCAUAUCAGAUU ...((((.............................................((((((.......))))))........(((....)))....)))).............. (-11.94 = -11.83 + -0.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:16:18 2011