
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,433,865 – 5,433,962 |
| Length | 97 |
| Max. P | 0.907427 |

| Location | 5,433,865 – 5,433,962 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 79.47 |
| Shannon entropy | 0.37092 |
| G+C content | 0.51822 |
| Mean single sequence MFE | -32.70 |
| Consensus MFE | -20.98 |
| Energy contribution | -23.66 |
| Covariance contribution | 2.68 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.903186 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
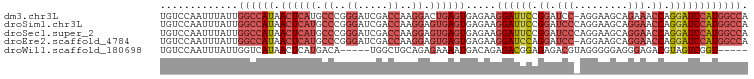
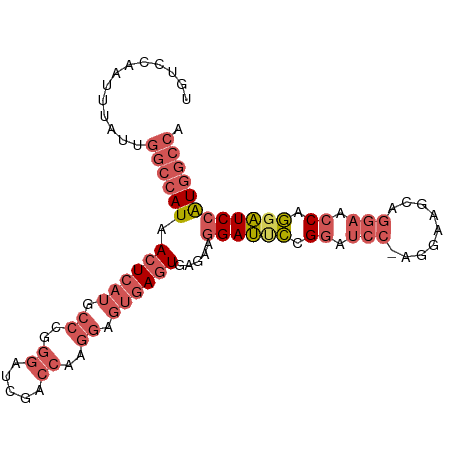
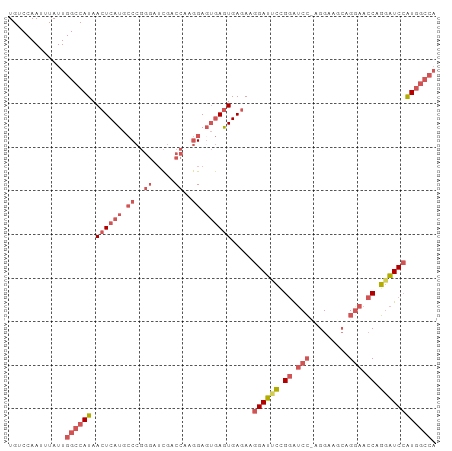
>dm3.chr3L 5433865 97 + 24543557 UGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGACUGAGUGAGAAGGAUUCCGGAUCC-AGGAAGCAGAAACCAGGAUCCAUGGCCA ............(((((((.(((((..((..((.....))..))..)))))............((((((-.((.........)).))))))))))))) ( -32.10, z-score = -1.52, R) >droSim1.chr3L 4950780 98 + 22553184 UGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUUCCGGAUCCCAGGAAGCAGGAACCAGGAUCCAUGGCCA ............(((((((.((((((.((..((.....))..)).)))))).....((((((.((.(((..(....).))).)).))))))))))))) ( -36.60, z-score = -2.31, R) >droSec1.super_2 5378990 98 + 7591821 UGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUUCCGGAUCCCAGGAAGCAGGAACCAGGAUCCAUGGCCA ............(((((((.((((((.((..((.....))..)).)))))).....((((((.((.(((..(....).))).)).))))))))))))) ( -36.60, z-score = -2.31, R) >droEre2.scaffold_4784 8127257 97 + 25762168 UGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUCCAGGAUCC-AGGAAGCAGGAACCAGGAUCCAUGGCCA ............(((((((.((((((.((..((.....))..)).)))))).....((((((.((.(((-.(....).))).)).))))))))))))) ( -39.80, z-score = -3.55, R) >droWil1.scaffold_180698 4200799 88 + 11422946 UGUCCAAUUUAUUGGUCAUAACUCAUGACA-----UGGCUGCAGAGAAAAUGACAGAGACGGAGAGACGUAGGGGGAGGGAGACGUAGUCGGU----- ...((.........(((((.....))))).-----.((((((...........(....(((......)))....)...(....))))))))).----- ( -18.40, z-score = -0.16, R) >consensus UGUCCAAUUUAUUGGCCAUAACUCAUGCCCGGGAUCGACCAAGGAGUGAGUGAGAAGGAUUCCGGAUCC_AGGAAGCAGGAACCAGGAUCCAUGGCCA .............((((((.((((((.((..((.....))..)).)))))).....((((((.((.(((.........))).)).)))))))))))). (-20.98 = -23.66 + 2.68)
| Location | 5,433,865 – 5,433,962 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 79.47 |
| Shannon entropy | 0.37092 |
| G+C content | 0.51822 |
| Mean single sequence MFE | -32.38 |
| Consensus MFE | -21.10 |
| Energy contribution | -23.22 |
| Covariance contribution | 2.12 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.907427 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
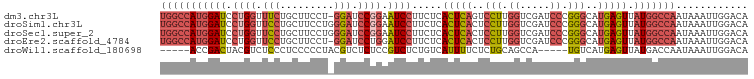
>dm3.chr3L 5433865 97 - 24543557 UGGCCAUGGAUCCUGGUUUCUGCUUCCU-GGAUCCGGAAUCCUUCUCACUCAGUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACA (((((((((((((.((.........)).-))))))............(((((((((..((.....)).)))).))))).)))))))............ ( -34.90, z-score = -1.67, R) >droSim1.chr3L 4950780 98 - 22553184 UGGCCAUGGAUCCUGGUUCCUGCUUCCUGGGAUCCGGAAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACA (((((((((((.((((.((((.......)))).)))).)))).....(((((..(((.((.....)).)))..))))).)))))))............ ( -37.20, z-score = -2.08, R) >droSec1.super_2 5378990 98 - 7591821 UGGCCAUGGAUCCUGGUUCCUGCUUCCUGGGAUCCGGAAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACA (((((((((((.((((.((((.......)))).)))).)))).....(((((..(((.((.....)).)))..))))).)))))))............ ( -37.20, z-score = -2.08, R) >droEre2.scaffold_4784 8127257 97 - 25762168 UGGCCAUGGAUCCUGGUUCCUGCUUCCU-GGAUCCUGGAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACA (((((((((((((.((.(((........-))).)).)))))).....(((((..(((.((.....)).)))..))))).)))))))............ ( -39.60, z-score = -3.27, R) >droWil1.scaffold_180698 4200799 88 - 11422946 -----ACCGACUACGUCUCCCUCCCCCUACGUCUCUCCGUCUCUGUCAUUUUCUCUGCAGCCA-----UGUCAUGAGUUAUGACCAAUAAAUUGGACA -----...(((...))).............((((..........(((((...(((((((....-----)).)).)))..))))).........)))). ( -13.01, z-score = -0.67, R) >consensus UGGCCAUGGAUCCUGGUUCCUGCUUCCU_GGAUCCGGAAUCCUUCUCACUCACUCCUUGGUCGAUCCCGGGCAUGAGUUAUGGCCAAUAAAUUGGACA (((((((((((.((((.(((.........))).)))).)))).....(((((..(((.((.....)).)))..))))).)))))))............ (-21.10 = -23.22 + 2.12)
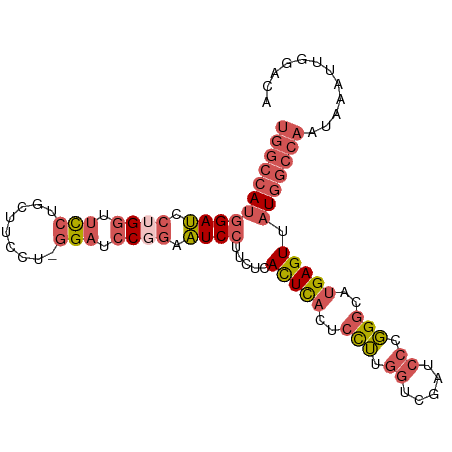
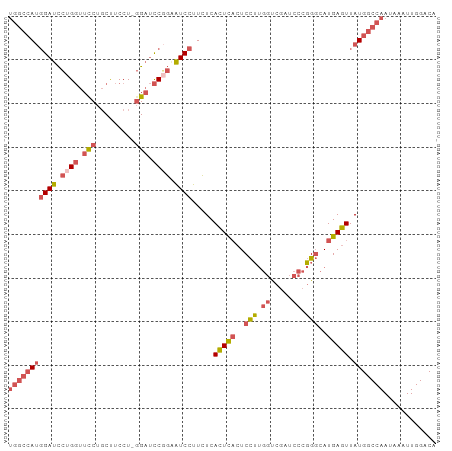
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:05:12 2011