| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,399,589 – 5,399,697 |
| Length | 108 |
| Max. P | 0.941561 |

| Location | 5,399,589 – 5,399,697 |
|---|---|
| Length | 108 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.27 |
| Shannon entropy | 0.49138 |
| G+C content | 0.36892 |
| Mean single sequence MFE | -13.52 |
| Consensus MFE | -10.60 |
| Energy contribution | -10.44 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.941561 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
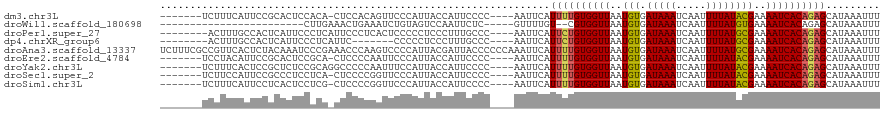
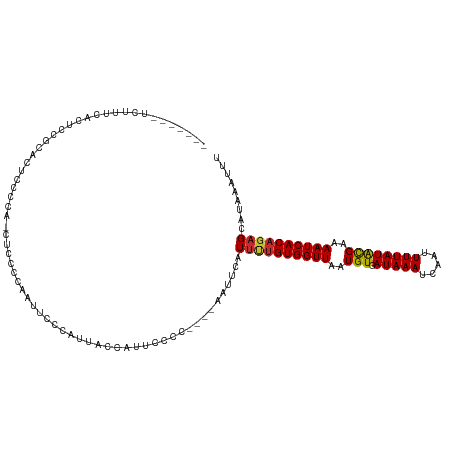
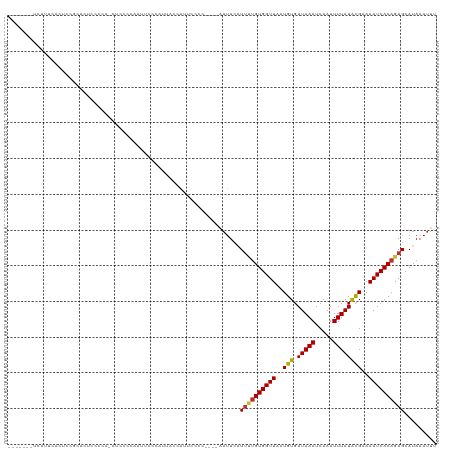
>dm3.chr3L 5399589 108 - 24543557 -------UCUUUCAUUCCGCACUCCACA-CUCCACAGUUCCCAUUACCAUUCCCC----AAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUACGAAAAUCACAGAGCAUAAAUUU -------...........((.......(-((....))).................----.......(((((((((..((((.((((.....))))))))..)))))))))))........ ( -12.00, z-score = -1.31, R) >droWil1.scaffold_180698 7181993 89 + 11422946 ------------------------CUUGAAACUGAAAUCUGUAGUCCAAUUCUC-----GUUUUGU--CGUGGUUAAUGUGAUAAAUCAAUUUUAUGUGAAAAUCACAGAGCAUAAAUUU ------------------------..(((((.(((.(((..(((.(((...(..-----.....).--..))))))....)))...))).)))))((((.....))))............ ( -9.40, z-score = 1.36, R) >droPer1.super_27 182601 108 - 1201981 --------ACUUUGCCACUCAUUCCCUCAUUCCCUCACUCCCCCUCCCUUUGCCC----AAUUCAUUCUGUGGUUAAUGUGAUAAAUCAAUUUUAUGCGAAAAUCACAGAGCAUAAAUUU --------...............................................----......((((((((((..(((.(((((.....))))))))..))))))))))......... ( -13.60, z-score = -1.46, R) >dp4.chrXR_group6 5060027 101 + 13314419 --------ACUUUGCCACUCAUUCCCUCAUUC-------CCCCCUCCCUUUGCCC----AAUUCAUUCUGUGGUUAAUGUGAUAAAUCAAUUUUAUGCGAAAAUCACAGAGCAUAAAUUU --------........................-------................----......((((((((((..(((.(((((.....))))))))..))))))))))......... ( -13.60, z-score = -1.41, R) >droAna3.scaffold_13337 10149557 120 + 23293914 UCUUUCGCCGUUCACUCUACAAAUCCCGAAACCCAAGUCCCCAUUACGAUUACCCCCCAAAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUGCGAAAAUCACAGAGCAUAAAUUU .........((((......................((((........)))).................(((((((..(((.(((((.....))))))))..)))))))))))........ ( -13.70, z-score = -0.40, R) >droEre2.scaffold_4784 8093084 108 - 25762168 -------UCCUACAUUCCGCACUCCGCA-CUCCCCAAUUCCCAUUACCAUUCCCC----AAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUACGAAAAUCACAGAGCAUAAAUUU -------...........((.....)).-..........................----......((((((((((..((((.((((.....))))))))..))))))))))......... ( -12.30, z-score = -1.61, R) >droYak2.chr3L 5966882 109 - 24197627 -------UCUUUCACUCCGCUCUCCGCAGGCCCCCAAUUUCCAUUACCAUUCCCC----AAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUACGAAAAUCACAGAGCAUAAAUUU -------...........((((......((...))....................----.........(((((((..((((.((((.....))))))))..)))))))))))........ ( -14.90, z-score = -1.27, R) >droSec1.super_2 5345327 108 - 7591821 -------UCUUCCAUUCCGCCCUCCUCA-CUCCCCGGUUCCCAUUACCAUUCCCC----AAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUACGAAAAUCACAGAGCAUAAAUUU -------...........((........-......(((.......))).......----.......(((((((((..((((.((((.....))))))))..)))))))))))........ ( -14.80, z-score = -1.75, R) >droSim1.chr3L 4915798 108 - 22553184 -------UCUUUCAUUCCUCACUCCUCG-CUCCCCGGUUCCCAUUACCAUUCCCC----AAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUACGAAAAUCACAGAGCAUAAAUUU -------....................(-(((...(((.......))).......----.........(((((((..((((.((((.....))))))))..)))))))))))........ ( -17.40, z-score = -2.82, R) >consensus _______UCUUUCACUCCGCACUCCCCA_CUCCCCAAUUCCCAUUACCAUUCCCC____AAUUCAUUUUGUGGUUAAUGUGAUAAAUCAAUUUUAUACGAAAAUCACAGAGCAUAAAUUU .................................................................((((((((((..(((.(((((.....))))))))..))))))))))......... (-10.60 = -10.44 + -0.16)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:05:00 2011