| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,389,118 – 5,389,208 |
| Length | 90 |
| Max. P | 0.902400 |

| Location | 5,389,118 – 5,389,208 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 74.39 |
| Shannon entropy | 0.50845 |
| G+C content | 0.49452 |
| Mean single sequence MFE | -22.22 |
| Consensus MFE | -13.52 |
| Energy contribution | -14.59 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.902400 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5389118 90 + 24543557 GGUUGGUCCGCUACCCACUUCCUUCGCCUGUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCACAGAAAUCCGGCCCC-------- ((.(((....))).)).............(((((((((((((......))))).....(((((.....)))))......))))))))...-------- ( -22.50, z-score = -1.54, R) >droSim1.chr3L 4905428 90 + 22553184 GGUUGGUCCCCUCCCCAUUUCCUUCGCCUGUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCACAGAAAUCCGGCCCC-------- ((.(((........)))...)).......(((((((((((((......))))).....(((((.....)))))......))))))))...-------- ( -22.20, z-score = -1.74, R) >droSec1.super_2 5334967 90 + 7591821 GGUUGGUCCCCUUUCCACAUCCUUCGCCUGUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCACAGAAAUCCGGCCCC-------- ((.(((........)))...)).......(((((((((((((......))))).....(((((.....)))))......))))))))...-------- ( -22.00, z-score = -1.62, R) >droYak2.chr3L 5956335 90 + 24197627 GGUUGGUUCCCUUACCACUUCCCUGGCCUGUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCACAGGAAUCCGGCCCC-------- ((((((((((............((((...(((((.....)))))...)))).......(((((.....)))))....)))).))))))..-------- ( -27.50, z-score = -2.17, R) >droEre2.scaffold_4784 8081262 90 + 25762168 GGUUGGUUCCCUUACCACUUCCUUGGCCUGUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCCCAGGAAUCCGGCCCC-------- ((((((((((....(((......)))......((..((((((......))))))....(((((.....))))).)).)))).))))))..-------- ( -25.40, z-score = -1.50, R) >droVir3.scaffold_13049 13935214 85 - 25233164 ----GGCUGGCUGGCUGCCUGC-UCGCCUGUCGCUGCGCCUGAUUUGUAAGAUAUUUUGCUAUUUUUUAUUGCCACACAAUUUCGGCUGU-------- ----(((.(((.(((.....))-).))).))).....(((.((.((((((((((......))))))..........)))).)).)))...-------- ( -19.40, z-score = 0.98, R) >droAna3.scaffold_13337 10139893 69 - 23293914 --------------------CCUUCGCC-GUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCCCAGGAAUCCGCCGGC-------- --------------------.....(((-(.(((((((((((......))))).....(((((.....)))))......)))))).))))-------- ( -23.20, z-score = -3.08, R) >dp4.chrXR_group6 8287011 88 + 13314419 ----------UUGUCUGCCAGCUCUAUCUGUCGGUUUGUCUGAUUUUCCAGAUAUUUUGCUAUUUUUUAUUGCCCCAGUAGCCCCCCUAGAGGACGCC ----------..........(((((.((((..((..((((((......))))))....((((((............)))))).))..))))))).)). ( -15.60, z-score = -0.29, R) >consensus GGUUGGUCCCCUUACCACUUCCUUCGCCUGUCGGAUUGUCUGAUUUCCCAGAUAUUUUGCUAUUUUUUAUGGCCACAGAAAUCCGGCCCC________ .............................(((((((((((((......))))).....(((((.....)))))......))))))))........... (-13.52 = -14.59 + 1.06)

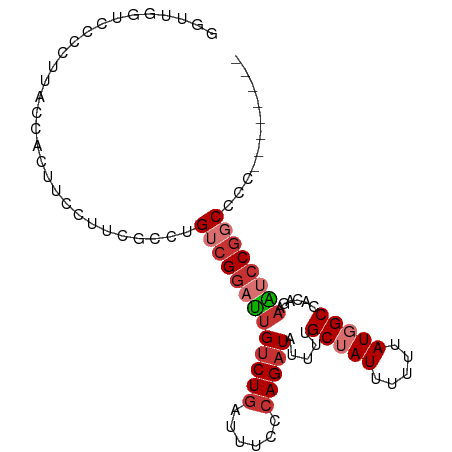

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:04:57 2011