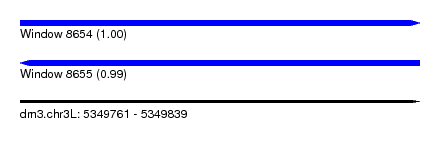
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,349,761 – 5,349,839 |
| Length | 78 |
| Max. P | 0.997292 |

| Location | 5,349,761 – 5,349,839 |
|---|---|
| Length | 78 |
| Sequences | 13 |
| Columns | 78 |
| Reading direction | forward |
| Mean pairwise identity | 95.51 |
| Shannon entropy | 0.11133 |
| G+C content | 0.53137 |
| Mean single sequence MFE | -27.95 |
| Consensus MFE | -27.90 |
| Energy contribution | -27.41 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.93 |
| Structure conservation index | 1.00 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.07 |
| SVM RNA-class probability | 0.997292 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5349761 78 + 24543557 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCUGUUA .((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))..... ( -27.10, z-score = -2.72, R) >droSim1.chr3L 4864783 78 + 22553184 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGUUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.09, R) >droSec1.super_2 5295020 78 + 7591821 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGUUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.09, R) >droYak2.chr3L 5916043 78 + 24197627 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGUUA (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.07, R) >droEre2.scaffold_4784 8041085 78 + 25762168 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGUUA (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.07, R) >droAna3.scaffold_13337 10097042 78 - 23293914 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGCUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.16, R) >dp4.chrXR_group6 5000838 78 - 13314419 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGCUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.16, R) >droPer1.super_27 123508 78 + 1201981 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGCUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.16, R) >droWil1.scaffold_180698 7115584 78 - 11422946 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGAUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.35, R) >droVir3.scaffold_13049 8121591 78 + 25233164 UGGUUCCAUCCGGGGUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGCUG (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -2.44, R) >droMoj3.scaffold_6680 11784336 78 + 24764193 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGUUA (((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))).... ( -28.30, z-score = -3.07, R) >droGri2.scaffold_15110 7150782 78 + 24565398 UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCCGCUG .((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))..... ( -26.70, z-score = -2.56, R) >anoGam1.chr2R 45370447 73 + 62725911 UGUUUCCGCCCGGGUUCGAACCGGGGACCUUCUGCGUGUGAAGCAGACGUGAUAACCACUACACUACGGAAAC----- .((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))----- ( -26.50, z-score = -2.16, R) >consensus UGGUUCCAUCCGGGAUCGAACCGGAGACCUUCUGCGUGUAAAGCAGACGUGAUAACCGCUACACUAUGGAACCAGUUG .((((((((((((.......))))).....(((((.......))))).(((.....))).......)))))))..... (-27.90 = -27.41 + -0.50)

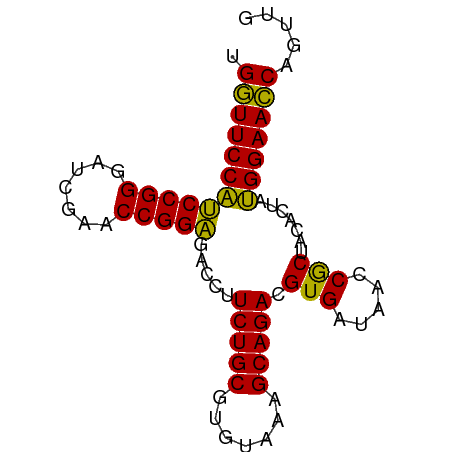
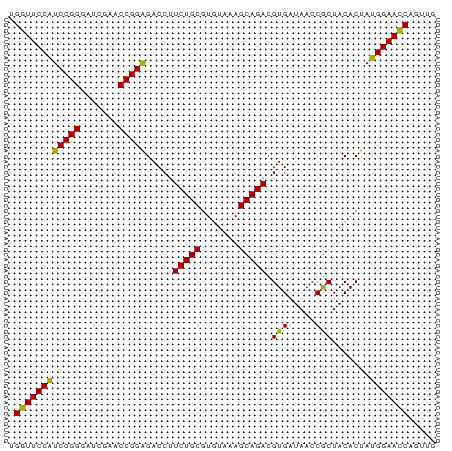
| Location | 5,349,761 – 5,349,839 |
|---|---|
| Length | 78 |
| Sequences | 13 |
| Columns | 78 |
| Reading direction | reverse |
| Mean pairwise identity | 95.51 |
| Shannon entropy | 0.11133 |
| G+C content | 0.53137 |
| Mean single sequence MFE | -26.99 |
| Consensus MFE | -27.39 |
| Energy contribution | -26.89 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.12 |
| Structure conservation index | 1.01 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.36 |
| SVM RNA-class probability | 0.989358 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5349761 78 - 24543557 UAACAGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.30, z-score = -2.61, R) >droSim1.chr3L 4864783 78 - 22553184 CAACUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.26, R) >droSec1.super_2 5295020 78 - 7591821 CAACUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.26, R) >droYak2.chr3L 5916043 78 - 24197627 UAACUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.28, R) >droEre2.scaffold_4784 8041085 78 - 25762168 UAACUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.28, R) >droAna3.scaffold_13337 10097042 78 + 23293914 CAGCUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.02, R) >dp4.chrXR_group6 5000838 78 + 13314419 CAGCUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.02, R) >droPer1.super_27 123508 78 - 1201981 CAGCUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.02, R) >droWil1.scaffold_180698 7115584 78 + 11422946 CAUCUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.29, R) >droVir3.scaffold_13049 8121591 78 - 25233164 CAGCUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGACCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -1.79, R) >droMoj3.scaffold_6680 11784336 78 - 24764193 UAACUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -2.28, R) >droGri2.scaffold_15110 7150782 78 - 24565398 CAGCGGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). ( -27.10, z-score = -1.80, R) >anoGam1.chr2R 45370447 73 - 62725911 -----GUUUCCGUAGUGUAGUGGUUAUCACGUCUGCUUCACACGCAGAAGGUCCCCGGUUCGAACCCGGGCGGAAACA -----(((((((.......(((.....))).(((((.......))))).....(((((.......)))))))))))). ( -25.50, z-score = -1.68, R) >consensus CAACUGGUUCCAUAGUGUAGCGGUUAUCACGUCUGCUUUACACGCAGAAGGUCUCCGGUUCGAUCCCGGAUGGAACCA .....(((((((..(((((((((.........)))))..))))..........(((((.......)))))))))))). (-27.39 = -26.89 + -0.50)
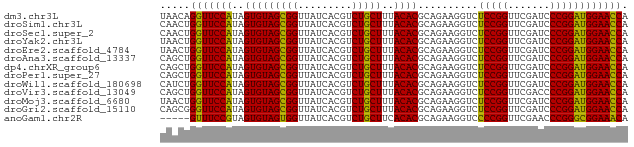
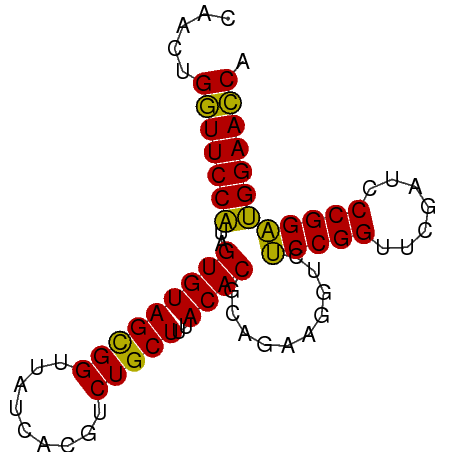
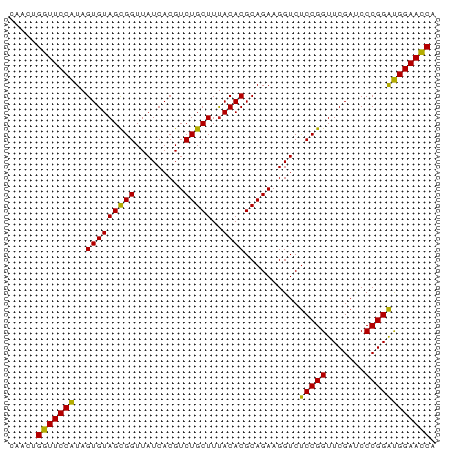
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:04:54 2011