
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,339,361 – 5,339,490 |
| Length | 129 |
| Max. P | 0.897800 |

| Location | 5,339,361 – 5,339,461 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 76.30 |
| Shannon entropy | 0.50155 |
| G+C content | 0.58455 |
| Mean single sequence MFE | -35.96 |
| Consensus MFE | -16.74 |
| Energy contribution | -16.61 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.804724 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5339361 100 + 24543557 AGGUGGUUCUGUGGAUGGU---UGGUGAA-GGUGCUGGAGGAGCAGGAGGUGCUGAUCUCGGUGGCCACGCUGUUGAGCGCCUUGGCCGAUUGGUUCAUGGGUG .......(((((((((.((---((((.((-(((((((((..((((.....))))..)))(((((....)))))...)))))))).))))).).))))))))).. ( -40.40, z-score = -2.69, R) >droSim1.chr3L 4853655 100 + 22553184 AGGUGGUUCUGUGGAUGGA---UGGUGAG-GGUGCUUGAGGAGCAGGAGGUGCUGAUCUCGGUGGCCACGCUGUUGAGCGCCUUGGCCGAUUGGUUCAUGGGUG .......(((((((((.((---((((.((-(((((((.((.(((.(..(.(((((....))))).)..)))).))))))))))).))).))).))))))))).. ( -41.50, z-score = -3.04, R) >droSec1.super_2 5284393 100 + 7591821 AGGUGGUUCUGUGGAUGGA---UGGUGAA-GGUUCUGGAGGAGCAGGAGGUGCUGAUCUCGGUGGCCACGCUGUUGAGCGCCUUGGCCAAUUGGUUCAUGGGUG .......(((((((((.((---((((...-.((((.....))))..((((((((.(...(((((....))))).).)))))))).))).))).))))))))).. ( -36.90, z-score = -1.69, R) >droYak2.chr3L 5905122 103 + 24197627 AGGUGGUUCUGUGGAUGGAAGAUGGUGAA-GGUGCUGGAGGAGCAGGAGGUGCUGAUCUCGGUGGCCACGCUGUUGAGCGCCUUGGCCGAUUGGUUCAUGGGUG .......(((((((((.((...((((.((-(((((((((..((((.....))))..)))(((((....)))))...)))))))).)))).)).))))))))).. ( -38.70, z-score = -2.24, R) >droEre2.scaffold_4784 8030501 100 + 25762168 AGGUGAUUCUGUGGAUGGA---UGGUGAA-GGUGCUGGAGGAGCAGGAGGUGCUGAUCUCGGUGGCCACGCUGUUGAGCGCCUUGGCCGUUUGGUUCAUGGGUG .......(((((((((.((---((((.((-(((((((((..((((.....))))..)))(((((....)))))...)))))))).)))).)).))))))))).. ( -40.40, z-score = -2.87, R) >droAna3.scaffold_13337 10086470 97 - 23293914 AGGUGGGUCAGUGGUUGGG---UGGUGCA-GGUGC---AGGAGCAGGAGGUGCUAAUCUUGGUGGCCACGCUGUUGAGCGCUUCGGCCGAGUGGUUCAUGGGUG ..(((..(((.(((((..(---(..(((.-...))---)...))..((((((((.....(((((....)))))...)))))))))))))..)))..)))..... ( -31.20, z-score = 0.37, R) >droPer1.super_27 113414 102 + 1201981 GGGGGAUGCAGGAGCUGGAUC-UGGAGCA-GGAGUAGGAGCAGGAGCAGGUGCUGUUGAUGGUGGCCACGCUGUAAAGCGAUGCGGCCGAGUGGUUCAUUGGUG ...((((.(((...))).)))-).((((.-..........(((.(((....))).))).((((.((..((((....))))..)).))))....))))....... ( -30.00, z-score = 0.41, R) >droWil1.scaffold_180698 7097230 82 - 11422946 ----------------------UGAUGAAUGAUGGUAAUCUGAUGGUCGUCAGUGAUCACGGUGGCCGCGCUAGAAAGCGUUUCGGCCAAAUGGUUCAUUGGAU ----------------------.(((((((.(((((...(((((....)))))..)))....((((((((((....))))...)))))).)).))))))).... ( -28.60, z-score = -2.29, R) >consensus AGGUGGUUCUGUGGAUGGA___UGGUGAA_GGUGCUGGAGGAGCAGGAGGUGCUGAUCUCGGUGGCCACGCUGUUGAGCGCCUUGGCCGAUUGGUUCAUGGGUG ........................................((((.......((((....))))(((((((((....))))...))))).....))))....... (-16.74 = -16.61 + -0.12)



| Location | 5,339,361 – 5,339,461 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 76.30 |
| Shannon entropy | 0.50155 |
| G+C content | 0.58455 |
| Mean single sequence MFE | -18.99 |
| Consensus MFE | -14.01 |
| Energy contribution | -13.64 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.897800 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
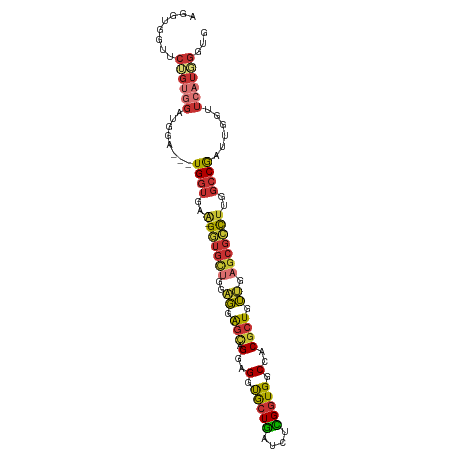
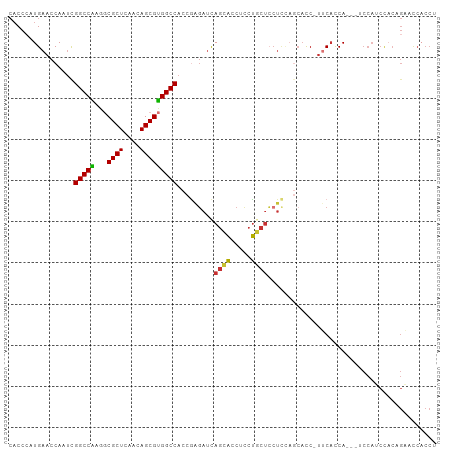
>dm3.chr3L 5339361 100 - 24543557 CACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC-UUCACCA---ACCAUCCACAGAACCACCU ......((((......((((...(((((....)))))))))...(((...((((.....)))).))).......-))))...---................... ( -18.80, z-score = -0.93, R) >droSim1.chr3L 4853655 100 - 22553184 CACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCAAGCACC-CUCACCA---UCCAUCCACAGAACCACCU ......(((.......((((...(((((....)))))))))...(((...((((.....)))).))).......-.)))...---................... ( -19.10, z-score = -1.00, R) >droSec1.super_2 5284393 100 - 7591821 CACCCAUGAACCAAUUGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGAACC-UUCACCA---UCCAUCCACAGAACCACCU ......((((.....(((((...(((((....))))))))))..(((...((((.....)))).))).......-))))...---................... ( -19.10, z-score = -0.86, R) >droYak2.chr3L 5905122 103 - 24197627 CACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC-UUCACCAUCUUCCAUCCACAGAACCACCU ......((((......((((...(((((....)))))))))...(((...((((.....)))).))).......-))))......................... ( -18.80, z-score = -0.78, R) >droEre2.scaffold_4784 8030501 100 - 25762168 CACCCAUGAACCAAACGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC-UUCACCA---UCCAUCCACAGAAUCACCU ......((((......((((...(((((....)))))))))...(((...((((.....)))).))).......-))))...---................... ( -18.80, z-score = -0.87, R) >droAna3.scaffold_13337 10086470 97 + 23293914 CACCCAUGAACCACUCGGCCGAAGCGCUCAACAGCGUGGCCACCAAGAUUAGCACCUCCUGCUCCU---GCACC-UGCACCA---CCCAACCACUGACCCACCU ................((((...(((((....)))))))))..........(((.....(((....---)))..-)))....---................... ( -17.40, z-score = -0.48, R) >droPer1.super_27 113414 102 - 1201981 CACCAAUGAACCACUCGGCCGCAUCGCUUUACAGCGUGGCCACCAUCAACAGCACCUGCUCCUGCUCCUACUCC-UGCUCCA-GAUCCAGCUCCUGCAUCCCCC ................((((((...(((....)))))))))..........(((...(((.(((...(......-.)...))-)....)))...)))....... ( -17.90, z-score = -0.13, R) >droWil1.scaffold_180698 7097230 82 + 11422946 AUCCAAUGAACCAUUUGGCCGAAACGCUUUCUAGCGCGGCCACCGUGAUCACUGACGACCAUCAGAUUACCAUCAUUCAUCA---------------------- ....(((((......((((((...((((....))))))))))..((((((..(((......)))))))))..))))).....---------------------- ( -22.00, z-score = -3.23, R) >consensus CACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC_UUCACCA___UCCAUCCACAGAACCACCU ................(((((...((((....))))))))).........((((.....))))......................................... (-14.01 = -13.64 + -0.37)


| Location | 5,339,387 – 5,339,490 |
|---|---|
| Length | 103 |
| Sequences | 8 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 76.72 |
| Shannon entropy | 0.49157 |
| G+C content | 0.57702 |
| Mean single sequence MFE | -26.95 |
| Consensus MFE | -18.32 |
| Energy contribution | -18.27 |
| Covariance contribution | -0.05 |
| Combinations/Pair | 1.36 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.787605 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
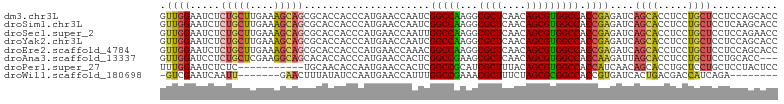
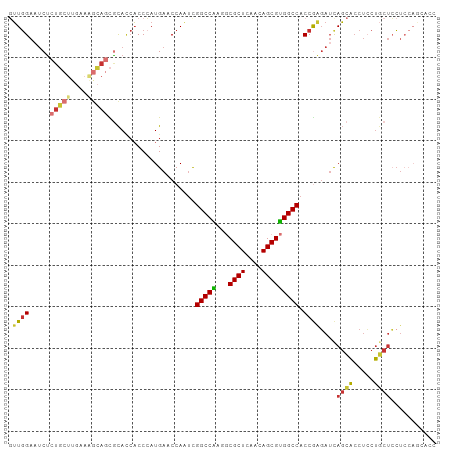
>dm3.chr3L 5339387 103 - 24543557 GUUGGAAUCUCUGCUUGAAAGCAGCGCACCACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC (((((((((((((((....))))......................((((...(((((....)))))))))...))))).((((.....))))..))))))... ( -31.20, z-score = -1.28, R) >droSim1.chr3L 4853681 103 - 22553184 GUUGGAAUCUCUGCUUGAAAGCAGCGCACCACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCAAGCACC ...........(((((((.(((((.....................((((...(((((....)))))))))...(((........))))))))..))))))).. ( -30.40, z-score = -1.06, R) >droSec1.super_2 5284419 103 - 7591821 GUUGGAAUCUCUGCUUGAAAGCAGCGCACCACCCAUGAACCAAUUGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGAACC .((((((((((((((....)))).....................(((((...(((((....))))))))))..))))).((((.....))))..))))).... ( -27.80, z-score = -0.39, R) >droYak2.chr3L 5905151 103 - 24197627 GUUGGAAUCUCUGCUUGAAAGCAGCGCACCACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC (((((((((((((((....))))......................((((...(((((....)))))))))...))))).((((.....))))..))))))... ( -31.20, z-score = -1.28, R) >droEre2.scaffold_4784 8030527 103 - 25762168 GUUGGAAUCUCUGCUUGAAAGCAGCGCACCACCCAUGAACCAAACGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC (((((((((((((((....))))......................((((...(((((....)))))))))...))))).((((.....))))..))))))... ( -31.20, z-score = -1.33, R) >droAna3.scaffold_13337 10086496 100 + 23293914 GUUGGAUCCUCUGCUCGAAGGCAGCACACCACCCAUGAACCACUCGGCCGAAGCGCUCAACAGCGUGGCCACCAAGAUUAGCACCUCCUGCUCCUGCACC--- ((.(((...(((((((....).)))....................((((...(((((....)))))))))....)))...(((.....)))))).))...--- ( -24.00, z-score = 0.50, R) >droPer1.super_27 113442 92 - 1201981 UUUGGAAUCUCUC-----------UGCAACACCAAUGAACCACUCGGCCGCAUCGCUUUACAGCGUGGCCACCAUCAACAGCACCUGCUCCUGCUCCUACUCC ...(((.......-----------(((.........((.....))((((((...(((....)))))))))..........)))...((....)))))...... ( -18.30, z-score = -0.96, R) >droWil1.scaffold_180698 7097246 87 + 11422946 -GUCGAAUCAAUU-------GAACUUUAUAUCCAAUGAACCAUUUGGCCGAAACGCUUUCUAGCGCGGCCACCGUGAUCACUGACGACCAUCAGA-------- -((((..((....-------))..........((.(((..(((.((((((...((((....))))))))))..))).))).)).)))).......-------- ( -21.50, z-score = -2.15, R) >consensus GUUGGAAUCUCUGCUUGAAAGCAGCGCACCACCCAUGAACCAAUCGGCCAAGGCGCUCAACAGCGUGGCCACCGAGAUCAGCACCUCCUGCUCCUCCAGCACC .((((.....(((((....))))).....................(((((...((((....))))))))).))))....((((.....))))........... (-18.32 = -18.27 + -0.05)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:04:47 2011