
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,170,801 – 5,170,893 |
| Length | 92 |
| Max. P | 0.747916 |

| Location | 5,170,801 – 5,170,893 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 67.62 |
| Shannon entropy | 0.58732 |
| G+C content | 0.49442 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -11.38 |
| Energy contribution | -13.19 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.747916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
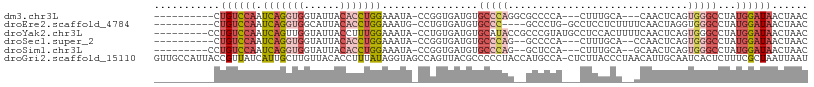
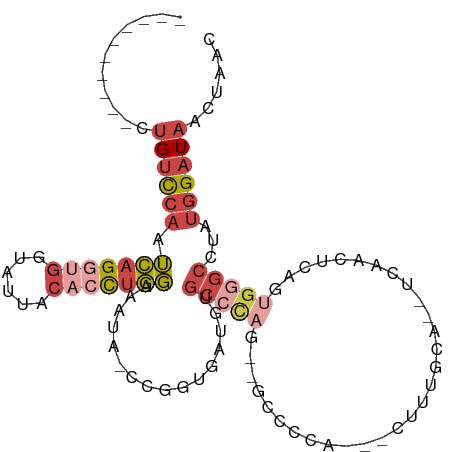
>dm3.chr3L 5170801 92 - 24543557 ----------CUGUCCAAUCAGGUGGUAUUACACCUGGAAAUA-CCGGUGAUGUGCCCAGGCGCCCCA---CUUUGCA---CAACUCAGUGGGCCUAUGGAUAACUAAC ----------.((((((.(((((((......))))))).....-..((....((((....))))((((---((..(..---...)..))))))))..))))))...... ( -31.90, z-score = -1.77, R) >droEre2.scaffold_4784 7863566 93 - 25762168 ----------CUGUCCAAUCAGGUGGCAUUACACCUGGAAAUG-CCUGUGAUGUGCCC----GCCCUG-GCCUCCUCUUUUCAACUAGGUGGGCCUAUGGAUAACUAAC ----------.((((((.(((((((......))))))).....-..........((((----(((.((-(..............))))))))))...))))))...... ( -31.14, z-score = -1.88, R) >droYak2.chr3L 5732122 99 - 24197627 ---------CCUGUCCAAUCAGUUGGUAUUACCUUUGGAAAUA-CCUGUGAUGUGCAUACCGCCCGUAUGCCUCCACUUUUCAACUCAGUGGGCCUAUGGAUAACUAAC ---------((.(((((...((((((((((.((...)).))))-)).(((..(.((((((.....)))))))..))).....))))...)))))....))......... ( -26.60, z-score = -1.91, R) >droSec1.super_2 5122402 91 - 7591821 ----------CUGUCCAAUCAGGUGGUAUUACACCUGGAAAUA-CCGGUGAUGUGCCCAG--GCCCCA---CUUUGCA--CCAACUCAGUGGGCCUAUGGAUAACUAAC ----------.((((((.(((((((......))))))).....-..(((.....))).((--((((.(---((..(..--....)..))))))))).))))))...... ( -32.10, z-score = -1.95, R) >droSim1.chr3L 4690646 92 - 22553184 ---------CCUGUCCAAUCAGGUGGUAUUACACCUGGAAAUA-CCGGUGAUGUGCCCAG--GCUCCA---CUUUGCA--GCAACUCAGUGGGCCUAUGGAUAACUAAC ---------..((((((.(((((((......))))))).....-..(((.....))).((--((.(((---((..(..--....)..))))))))).))))))...... ( -32.10, z-score = -1.89, R) >droGri2.scaffold_15110 6949916 108 - 24565398 GUUGCCAUUACCGUUAUCAUUGCUUGUUACACCUUUAUAGGUAGCCAGUUACGCCCCCUACCAUGCCA-CUCUUACCCUAACAUUGCAAUCACUCUUUCGCUAAUUAAU ......((((.((.....(((((.(((((.((((....)))).((.......))..............-.........)))))..)))))........)).)))).... ( -10.42, z-score = 0.41, R) >consensus __________CUGUCCAAUCAGGUGGUAUUACACCUGGAAAUA_CCGGUGAUGUGCCCAG__GCCCCA___CUUUGCA__UCAACUCAGUGGGCCUAUGGAUAACUAAC ...........((((((.(((((((......)))))))................(((((..............................)))))...))))))...... (-11.38 = -13.19 + 1.81)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:04:25 2011