
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,164,074 – 5,164,182 |
| Length | 108 |
| Max. P | 0.657284 |

| Location | 5,164,074 – 5,164,180 |
|---|---|
| Length | 106 |
| Sequences | 10 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 66.39 |
| Shannon entropy | 0.70440 |
| G+C content | 0.49474 |
| Mean single sequence MFE | -28.28 |
| Consensus MFE | -10.04 |
| Energy contribution | -10.04 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.657284 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5164074 106 + 24543557 -------GCAGAAAGAGAGCGGUAGUGUGGCAGAUGCACGGGAAGAAAGAGACAGGCACAUUGGCAUGCUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA -------((.........)).....(((((((......(.....)...((((.(.((.(((((.((((((.......)))))--).))))))).).))))........))))))) ( -28.90, z-score = -1.17, R) >droSim1.chr3L 4683755 106 + 22553184 -------GCAGAAAGAGAGCGGUAGUGAGGCAGAUGCACGGGAAGAAAGAGACAGCCACAUUGGCAUGCUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA -------...........(.((((.((((((....)).......((((((....((((..(((.((((((.......)))))--).))))))).)))))).))))...)))).). ( -28.20, z-score = -1.24, R) >droSec1.super_2 5115783 106 + 7591821 -------GCAGAAAGAGAGCGGUAGUGAGGCAGAUGCACGGGAAGAAAGAGACAGCCACAUUGGCAUGCUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA -------...........(.((((.((((((....)).......((((((....((((..(((.((((((.......)))))--).))))))).)))))).))))...)))).). ( -28.20, z-score = -1.24, R) >droYak2.chr3L 5725446 106 + 24197627 -------GCAGAAAGAGAGCGGUAGAGAGGCAGAUGCACGGGAAGAAAGAGACAGGCACAUUGGCAUGUUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA -------...........(.((((..(((((....))...........((((.(.((.(((((.((((((.......)))))--).))))))).).)))).)))....)))).). ( -24.80, z-score = -1.10, R) >droEre2.scaffold_4784 7856800 106 + 25762168 -------GCAGAAAGAGAGCGGUGGAGAGGUAGCUGCACGGUCAGAAAGAGACAGGCACAUUGGCAUGUUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA -------(((....(((.((.(((.((......)).))).))......((((.(.((.(((((.((((((.......)))))--).))))))).).)))).)))....))).... ( -28.30, z-score = -0.83, R) >droAna3.scaffold_13337 9920160 88 - 23293914 --------------------------GCAGAAGGAGCUUCGG-AUAGUGCCACAGAGGCAGUGGCUCGUUGACGACAGCCUCUCGACGAUGGCGUUUUUCACUUAAAAUGCCACA --------------------------(..((..(.((((((.-...(.(((((.......))))).).....))).))))..))..)(.(((((((((......)))))))))). ( -28.50, z-score = -1.16, R) >dp4.chrXR_group6 7399350 103 + 13314419 GGAUAGGUGGCAGUAG--GAUGCAUGCCCCAAGAGGGAGACAUAGAGAGACAU--------UGGCAUGUUGACGACAGCCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA ......((((((.(((--(....(((((((....))).).))).(((((((..--------.....((((...))))(((((--....)))))))))))).))))...)))))). ( -33.80, z-score = -3.11, R) >droPer1.super_153 92581 103 + 195207 GGAUAGGUGGCAGUAG--GAUGCAUGCCCCAAGAGGGAGACAUAGAGAGACAU--------UGGCAUGUUGACGACAGCCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACA ......((((((.(((--(....(((((((....))).).))).(((((((..--------.....((((...))))(((((--....)))))))))))).))))...)))))). ( -33.80, z-score = -3.11, R) >droMoj3.scaffold_6680 11538774 92 + 24764193 ------UUAGGUGUAGCUGAGGUACAGUUGGCAAAGGAUACACAGAA--------------AGGCAUGUGGACCAAA-CUGU--GAUGAUGGCGUUUUUCACUUAAAAUGCCACA ------.....((((((((.....))))).))).......(((((..--------------.((........))...-))))--)....(((((((((......))))))))).. ( -22.00, z-score = -0.09, R) >droVir3.scaffold_13049 7886839 111 + 25233164 --GCAUGUGUGGGUAGUCGAGGUAUAGUUGGCAAAGGAUACACUGAAGGAUAUACAGCGAAAGGCAUGUGGACCAAAGCUGU--GUUGAUGGCGUUUUUCACUUAAAAUGCCACA --.((.(((((....(((((.......)))))......)))))))......(((((((....((........))...)))))--))...(((((((((......))))))))).. ( -26.30, z-score = 0.52, R) >consensus _______GCAGAAAGAGAGCGGUAGAGAGGCAGAUGCACGGGAAGAAAGAGACAG_CACAUUGGCAUGUUGACGACAGCCAU__GACGAUGGCGUUUUUCACUUAAAAUGCCACA .......................................................................................(.(((((((((......)))))))))). (-10.04 = -10.04 + 0.00)



| Location | 5,164,076 – 5,164,182 |
|---|---|
| Length | 106 |
| Sequences | 10 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 69.40 |
| Shannon entropy | 0.62873 |
| G+C content | 0.48710 |
| Mean single sequence MFE | -28.58 |
| Consensus MFE | -10.04 |
| Energy contribution | -10.04 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.645420 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 5164076 106 + 24543557 -AGAAAGAGAGCGGUAGUGUGGCAGAUGCACGGGAAGAAAGAGACAGGCACAU------UGGCAUGCUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA -...............((((((((......(.....)...((((.(.((.(((------((.((((((.......)))))--).))))))).).))))........)))))))). ( -33.10, z-score = -2.97, R) >droSim1.chr3L 4683757 106 + 22553184 -AGAAAGAGAGCGGUAGUGAGGCAGAUGCACGGGAAGAAAGAGACAGCCACAU------UGGCAUGCUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA -...............(((.((((..........(((..((((((.((((..(------((.((((((.......)))))--).)))))))))))))..)))....)))).))). ( -28.44, z-score = -1.67, R) >droSec1.super_2 5115785 106 + 7591821 -AGAAAGAGAGCGGUAGUGAGGCAGAUGCACGGGAAGAAAGAGACAGCCACAU------UGGCAUGCUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA -...............(((.((((..........(((..((((((.((((..(------((.((((((.......)))))--).)))))))))))))..)))....)))).))). ( -28.44, z-score = -1.67, R) >droYak2.chr3L 5725448 106 + 24197627 -AGAAAGAGAGCGGUAGAGAGGCAGAUGCACGGGAAGAAAGAGACAGGCACAU------UGGCAUGUUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA -.........(.((((..(((((....))...........((((.(.((.(((------((.((((((.......)))))--).))))))).).)))).)))....)))).)... ( -24.80, z-score = -1.48, R) >droEre2.scaffold_4784 7856802 106 + 25762168 -AGAAAGAGAGCGGUGGAGAGGUAGCUGCACGGUCAGAAAGAGACAGGCACAU------UGGCAUGUUGACGACAGGCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA -.........((.(((.((......)).))).(((.......)))..))..((------((.((((((.......)))))--).))))((((((((......))))))))..... ( -25.70, z-score = -0.26, R) >droAna3.scaffold_13337 9920162 88 - 23293914 ---------------AGAAGGAGCUUCGGAUAGUGCCACA----GAGGCAG--------UGGCUCGUUGACGACAGCCUCUCGACGAUGGCGUUUUUCACUUAAAAUGCCACACA ---------------.((..(.((((((....(.(((((.----......)--------)))).).....))).))))..))...(.(((((((((......))))))))))... ( -28.30, z-score = -1.43, R) >dp4.chrXR_group6 7399350 105 + 13314419 GGAUAGGUGGCAGUAGGAUGCAUGCCCCAAGAGGGAGACAUAGAGAGACAU--------UGGCAUGUUGACGACAGCCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA ......((((((.((((....(((((((....))).).))).(((((((..--------.....((((...))))(((((--....)))))))))))).))))...))))))... ( -33.80, z-score = -2.98, R) >droPer1.super_153 92581 105 + 195207 GGAUAGGUGGCAGUAGGAUGCAUGCCCCAAGAGGGAGACAUAGAGAGACAU--------UGGCAUGUUGACGACAGCCAU--GACGAUGGCGUUUUUCACUUAAAAUGCCACACA ......((((((.((((....(((((((....))).).))).(((((((..--------.....((((...))))(((((--....)))))))))))).))))...))))))... ( -33.80, z-score = -2.98, R) >droMoj3.scaffold_6680 11538776 92 + 24764193 ------AGGUGUAGCUGAGGUACAGUUGGCAAAGGAUACACAGAA--------------AGGCAUGUGGACCAAA-CUGU--GAUGAUGGCGUUUUUCACUUAAAAUGCCACAAA ------...((((((((.....))))).))).......(((((..--------------.((........))...-))))--).((.(((((((((......))))))))))).. ( -22.80, z-score = -0.40, R) >droVir3.scaffold_13049 7886841 111 + 25233164 --AUGUGUGGGUAGUCGAGGUAUAGUUGGCAAAGGAUACACUGAAGGAUAUACAGCGAAAGGCAUGUGGACCAAAGCUGU--GUUGAUGGCGUUUUUCACUUAAAAUGCCACAAA --..(((((....(((((.......)))))......))))).........((((((....((........))...)))))--)(((.(((((((((......)))))))))))). ( -26.60, z-score = -0.09, R) >consensus _AGAAAGAGAGCGGUAGAGGGACAGAUGCACAGGGAGAAAGAGACAGGCACAU______UGGCAUGUUGACGACAGCCAU__GACGAUGGCGUUUUUCACUUAAAAUGCCACACA .....................................................................................(.(((((((((......))))))))))... (-10.04 = -10.04 + 0.00)

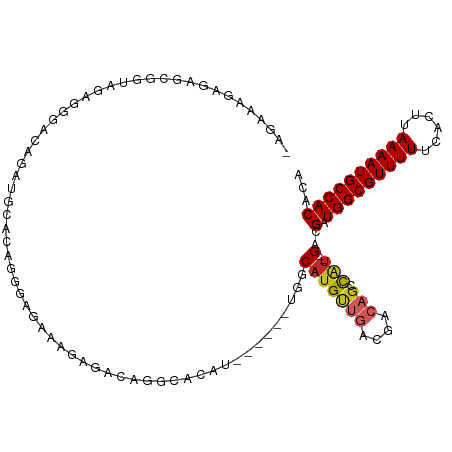

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:04:24 2011