
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 5,088,731 – 5,088,820 |
| Length | 89 |
| Max. P | 0.732518 |

| Location | 5,088,731 – 5,088,820 |
|---|---|
| Length | 89 |
| Sequences | 11 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 82.36 |
| Shannon entropy | 0.38605 |
| G+C content | 0.44530 |
| Mean single sequence MFE | -22.34 |
| Consensus MFE | -17.37 |
| Energy contribution | -17.29 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.732518 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
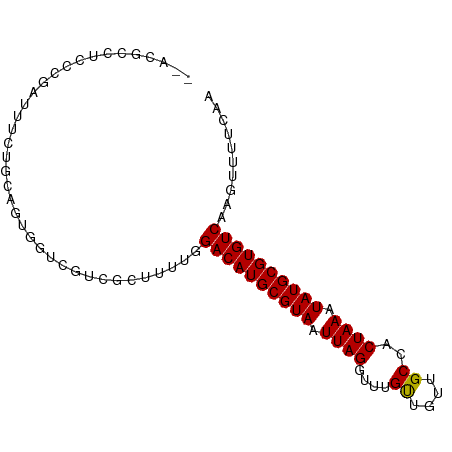
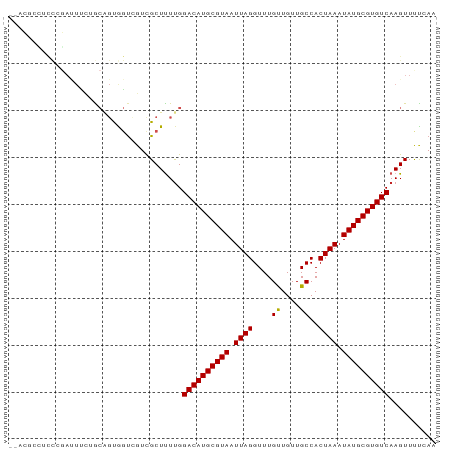
>dm3.chr3L 5088731 89 + 24543557 --ACUCCUCCCGAUUUCUGCAGUGGUCGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUUCAA --........((((..(....)..)))).........((((((((((.((((....((....))..)))).)))))))))).......... ( -20.20, z-score = -0.81, R) >droEre2.scaffold_4784 7791772 89 + 25762168 --ACGCCCCCCGAUUUCUGCAGUGGUCGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUUCAA --..((....((((..(....)..))))..)).....((((((((((.((((....((....))..)))).)))))))))).......... ( -21.20, z-score = -0.60, R) >droYak2.chr3L 5662028 89 + 24197627 --ACGCCUCCCGAUUUCUGCAGUGGUCGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUUCAA --..((....((((..(....)..))))..)).....((((((((((.((((....((....))..)))).)))))))))).......... ( -21.20, z-score = -0.54, R) >droSec1.super_2 5054584 89 + 7591821 --ACGCCUCCCGAUUUCUGCAGUGGUCGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUUCAA --..((....((((..(....)..))))..)).....((((((((((.((((....((....))..)))).)))))))))).......... ( -21.20, z-score = -0.54, R) >droSim1.chr3L 4623619 89 + 22553184 --ACGCCUCCCGAUUUCUGCAGUGGUCGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUUCAA --..((....((((..(....)..))))..)).....((((((((((.((((....((....))..)))).)))))))))).......... ( -21.20, z-score = -0.54, R) >dp4.chrXR_group3a 1214908 75 + 1468910 ---------------GCUG-GUCGCUGGUCGCUCUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUCCAA ---------------((..-...))(((..(((....((((((((((.((((....((....))..)))).)))))))))).)))..))). ( -22.20, z-score = -1.81, R) >droPer1.super_23 1234675 75 + 1662726 ---------------GCUG-GUCGCUGGUCGCUCUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUCCAA ---------------((..-...))(((..(((....((((((((((.((((....((....))..)))).)))))))))).)))..))). ( -22.20, z-score = -1.81, R) >droAna3.scaffold_13337 9855089 91 - 23293914 AUUUGACACCAGAUUUCUGCAGUGGUCGUUGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUCAAA ....((((((((....)))..)).)))(..((((...((((((((((.((((....((....))..)))).))))))))))))))..)... ( -23.60, z-score = -1.17, R) >droVir3.scaffold_13049 7820536 88 + 25233164 --ACUCGACUCGACUCG-ACUGCAGUGGUCGCUUUCGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUCUAAA --....(((((((..((-(((.....)))))...)))((((((((((.((((....((....))..)))).)))))))))).))))..... ( -25.30, z-score = -1.51, R) >droMoj3.scaffold_6680 11471436 89 + 24764193 --ACUCGAUCCCACUCGGACUGCAGCACUCGCUUUUGGACAUGCGUAAUUAGGUUUGCUGUUGCCACUAAAUAUGCGUGUCAAGUUCUAAA --..............(((((..(((....)))....((((((((((.((((....((....))..)))).)))))))))).))))).... ( -25.00, z-score = -1.80, R) >droGri2.scaffold_15110 6873172 86 + 24565398 ----ACUGUUGGAAU-GAACUGCAGUGGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUCUAAA ----....(((((((-.....((.......)).....((((((((((.((((....((....))..)))).))))))))))..))))))). ( -22.40, z-score = -1.11, R) >consensus __ACGCCUCCCGAUUUCUGCAGUGGUCGUCGCUUUUGGACAUGCGUAAUUAGGUUUGUUGUUGCCACUAAAUAUGCGUGUCAAGUUUUCAA .....................................((((((((((.((((....((....))..)))).)))))))))).......... (-17.37 = -17.29 + -0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:04:14 2011