
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 4,710,542 – 4,710,659 |
| Length | 117 |
| Max. P | 0.756875 |

| Location | 4,710,542 – 4,710,655 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.31 |
| Shannon entropy | 0.22098 |
| G+C content | 0.35310 |
| Mean single sequence MFE | -25.86 |
| Consensus MFE | -19.94 |
| Energy contribution | -19.30 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.643971 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
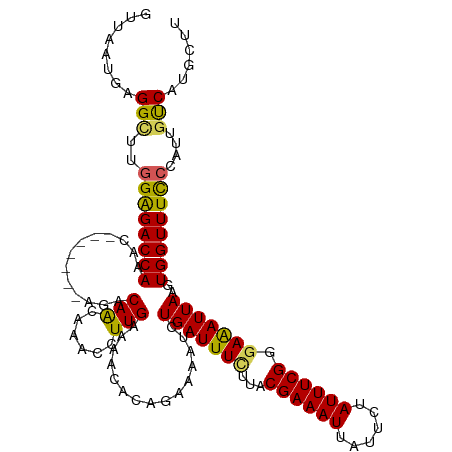
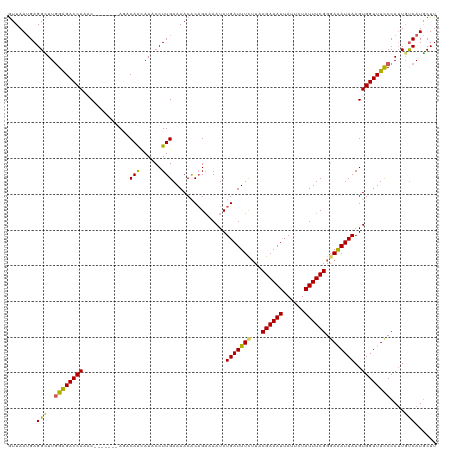
>dm3.chr3L 4710542 113 - 24543557 GUUAAUGAGGCUUGGAGACCAAAC-------AGACAACAAGCCUUGAAAACACAGAAAAUCUGAUUUCUUACGAAAUGAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAUUGUCAUGCUU (((..(((((((((..........-------......)))))))))..))).(((.....))).........((((((....))))))((((((((....))))))))............ ( -27.39, z-score = -1.75, R) >droSim1.chr3L 4242450 113 - 22553184 GUUAAUGAGGCUUUGAGACCAAAC-------AGACAACAAACCUUGAAAACACAGAAAAUCUGAUUUCUUACGAAAUUAUUCUAUUUCGGUAGAUUAAGUGGUUUUCCAUUGUCAUGCUU .......((((((((....)))).-------.(((((.....(((((.....(((.....)))...(((..((((((......))))))..))))))))(((....))))))))..)))) ( -20.60, z-score = -0.40, R) >droSec1.super_2 4694451 113 - 7591821 GUUAAUGAGGCUUGGGGACCAAGC-------AGACAACAAACCUUGAAAACACAGAAAAUCUGAUUUCAUACGAAAUUAUUAUAUUUCGGGAGAUUAAGUGGUUUUCCAUUCUCAUGCUU ....(((((((((((...))))))-------.......................((((((((((((((...((((((......)))))).)))))))...)))))))....))))).... ( -28.40, z-score = -2.49, R) >droYak2.chr3L 5292416 120 - 24197627 AUUAAUGAGGUUUGGAGACCAAACACAGCAAACACAGUAAACCUUGAAAACACCGAAAAUCUGAUUUCUUACGAAAUUAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAAUGCCAUGCUU ........(((..((((((((......((.......))....(((((.....((((((...(((((((....))))))).....))))))....)))))))))))))....)))...... ( -26.70, z-score = -1.88, R) >droEre2.scaffold_4784 7426371 110 - 25762168 GUUAAAGAGGUUUGGAGACCA--C-------AUACAA-AAACCUUGAAAACGCAGUGAAUCUGAUUUUUUACGAAAUUAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAUUGUCAUACUU ......((.(...((((((((--(-------......-......((((((..(((.....)))..))))))((((((......)))))).........)))))))))...).))...... ( -26.20, z-score = -2.08, R) >consensus GUUAAUGAGGCUUGGAGACCAAAC_______AGACAACAAACCUUGAAAACACAGAAAAUCUGAUUUCUUACGAAAUUAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAUUGUCAUGCUU ........(((..((((((((.............(((......)))...............(((((((...((((((......)))))).)))))))..))))))))....)))...... (-19.94 = -19.30 + -0.64)

| Location | 4,710,546 – 4,710,659 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.65 |
| Shannon entropy | 0.21497 |
| G+C content | 0.33733 |
| Mean single sequence MFE | -25.46 |
| Consensus MFE | -19.94 |
| Energy contribution | -19.30 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.756875 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
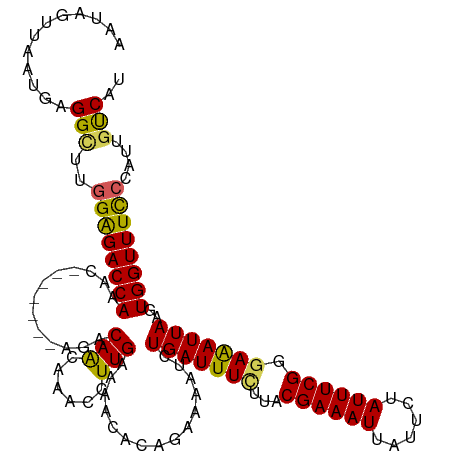
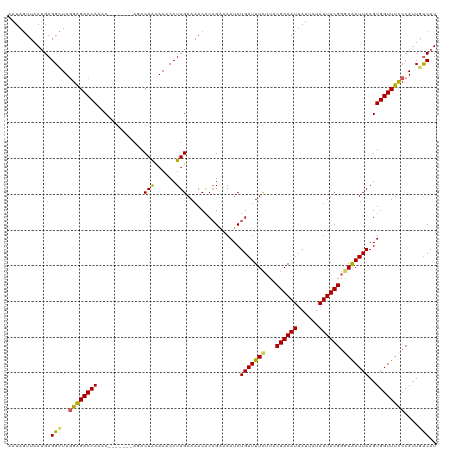
>dm3.chr3L 4710546 113 - 24543557 AAUAGUUAAUGAGGCUUGGAGACCAAAC-------AGACAACAAGCCUUGAAAACACAGAAAAUCUGAUUUCUUACGAAAUGAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAUUGUCAU ....(((..(((((((((..........-------......)))))))))..))).(((.....))).........((((((....))))))((((((((....))))))))........ ( -27.59, z-score = -2.10, R) >droSim1.chr3L 4242454 113 - 22553184 AAUAGUUAAUGAGGCUUUGAGACCAAAC-------AGACAACAAACCUUGAAAACACAGAAAAUCUGAUUUCUUACGAAAUUAUUCUAUUUCGGUAGAUUAAGUGGUUUUCCAUUGUCAU ............(((..((((((((...-------.....................(((.....)))...(((..((((((......))))))..))).....))))))..))..))).. ( -19.10, z-score = -0.13, R) >droSec1.super_2 4694455 113 - 7591821 AAUAGUUAAUGAGGCUUGGGGACCAAGC-------AGACAACAAACCUUGAAAACACAGAAAAUCUGAUUUCAUACGAAAUUAUUAUAUUUCGGGAGAUUAAGUGGUUUUCCAUUCUCAU ........(((((((((((...))))))-------.......................((((((((((((((...((((((......)))))).)))))))...)))))))....))))) ( -27.70, z-score = -2.60, R) >droYak2.chr3L 5292420 120 - 24197627 AAUAAUUAAUGAGGUUUGGAGACCAAACACAGCAAACACAGUAAACCUUGAAAACACCGAAAAUCUGAUUUCUUACGAAAUUAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAAUGCCAU ............(((..((((((((......((.......))....(((((.....((((((...(((((((....))))))).....))))))....)))))))))))))....))).. ( -26.70, z-score = -2.30, R) >droEre2.scaffold_4784 7426375 110 - 25762168 AAUAGUUAAAGAGGUUUGGAGACCA--C-------AUACAA-AAACCUUGAAAACGCAGUGAAUCUGAUUUUUUACGAAAUUAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAUUGUCAU ..........((.(...((((((((--(-------......-......((((((..(((.....)))..))))))((((((......)))))).........)))))))))...).)).. ( -26.20, z-score = -2.13, R) >consensus AAUAGUUAAUGAGGCUUGGAGACCAAAC_______AGACAACAAACCUUGAAAACACAGAAAAUCUGAUUUCUUACGAAAUUAUUCUAUUUCGGGAAAUUAAGUGGUUUCCCAUUGUCAU ............(((..((((((((.............(((......)))...............(((((((...((((((......)))))).)))))))..))))))))....))).. (-19.94 = -19.30 + -0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:03:30 2011