
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 4,481,372 – 4,481,461 |
| Length | 89 |
| Max. P | 0.863926 |

| Location | 4,481,372 – 4,481,461 |
|---|---|
| Length | 89 |
| Sequences | 11 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 65.69 |
| Shannon entropy | 0.71302 |
| G+C content | 0.35659 |
| Mean single sequence MFE | -18.35 |
| Consensus MFE | -7.18 |
| Energy contribution | -6.48 |
| Covariance contribution | -0.70 |
| Combinations/Pair | 2.00 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.863926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 4481372 89 + 24543557 UAAAUCAU-CCGAUAAAGGCUUCACUAGGUGAUCGAAUUGUGCUC--AUUUUUGAGUGCAGACCCUAC------UAUUAGAAUACCUCUUUUUCAUUC ........-..((.((((........(((((......(((..(((--(....))))..)))...(((.------...)))..))))))))).)).... ( -15.90, z-score = -0.66, R) >droSec1.super_2 4463982 89 + 7591821 UAAAUCAU-CCGAUAAAGGCUUCACUUGGUGAUUGAAUCGUGCUC--AUUUUGGAGUGCAGACCCUAC------UAUUAGAAUACCUCUUUUUCAUUC ..((((((-(.....(((......))))))))))(((..(..(((--......)))..).....(((.------...)))...........))).... ( -12.90, z-score = 0.82, R) >droYak2.chr3L 5053400 89 + 24197627 UAAAUCAU-CCGAUAAAGGCUUCACUAGGUGAUCGAUUUGUGCUC--AUUUUGGAGUGCAGACCCUAC------UAUUAGAAUACCUCCUUUUCAUUC ........-..((.(((((.......(((((.....((((..(((--......)))..))))..(((.------...)))..)))))))))))).... ( -17.31, z-score = -0.68, R) >droEre2.scaffold_4784 7187015 89 + 25762168 UAAAUCAU-CCGAUAAAGGCUUCACUAGGUGAUCGAUUUGUGCUU--AUUUUGGAGUGCAGACCCUAC------UAUUAGAAUACCUCCUUUUCAUUC ........-..((.(((((.......(((((.....((((..(((--......)))..))))..(((.------...)))..)))))))))))).... ( -15.41, z-score = -0.11, R) >droAna3.scaffold_13337 2561232 86 + 23293914 UAAACAUU-GCGAUUACGACUUCACUAGGUGAUCGAUUUGUGCUC--AUUUUGGAGUGCAGUCUUUAC------UAUUAGCAUACC---UUUUCAUUC .......(-((((((((...........))))))((((.(..(((--......)))..))))).....------.....)))....---......... ( -17.70, z-score = -1.41, R) >dp4.chrXR_group8 2576614 88 + 9212921 UAAACCAU-GCGAGAUCUGUUCUACUAAGUGAUCGAGUUGUACUC--AUUUUGGAGUGCAAACUUUA-------UAAUAAGAAUUUUUUUUUGCAUUC ......((-((((((...(((((.....((((.....((((((((--......))))))))...)))-------)....)))))....)))))))).. ( -20.90, z-score = -2.57, R) >droPer1.super_24 1084039 91 - 1556852 UAGACCCUUGCCAGAUCUCUUCUA-UAAGAGAUCGAGUUGUACUCCUGUAUGGAGGCGCGAACUUUAA------UAGGAGAAAUUUUUUUUUGCAUUC ........(((..((((((((...-.))))))))(((((...(((((((.(((((.......))))))------)))))).)))))......)))... ( -24.30, z-score = -1.98, R) >droWil1.scaffold_180698 592357 93 - 11422946 -UAAACAUAGCCGGAUUUGUUUUAUUCCGUGAUCGAUUUGUACUC--AUUUUGGAGUGCAGACUUUAUUUUAAG--AUUACAAUUCCACUUUCCACCC -...........(((.((((...(((....(((.(((((((((((--......))))))))).)).)))....)--)).)))).)))........... ( -17.90, z-score = -2.11, R) >droVir3.scaffold_12758 48902 93 + 972128 UAAAUCCUCAUCUGAAAUGAUUCACUAAGUGAUCGACUUGUACUC--AUUUUGGAGUGCAAGUUAAAUA-UACAUAAGAAUAUUUCCUCUUUUCGU-- .............((((((..((((...))))..(((((((((((--......))))))))))).....-..........))))))..........-- ( -19.70, z-score = -2.39, R) >droMoj3.scaffold_6680 1173614 94 + 24764193 UAAAGCCUCAUCUGAAAUGUUUCACUAAGUGAUCAACUUGUGCUC--AUUUUGGAGUGCAAGUUAAAUAAUAUGUAAGAAAAUUUCAUUUUUUCGC-- ..........(((...(((((((((...))))..((((((..(((--......)))..))))))....)))))...))).................-- ( -18.90, z-score = -1.60, R) >droGri2.scaffold_15110 23256912 93 - 24565398 -UAAGCCUCAUCUGAAAUGAUUCACUAAGUGAUCGGCUCGUACUC--AUUUUGGAGUAUGAGUUUAAUAUACAUUUAGAAUAUAUCCUCUUUUCGU-- -.........((((((..(((.(((...))))))(((((((((((--......))))))))))).........)))))).................-- ( -20.90, z-score = -2.02, R) >consensus UAAAUCAU_CCGAGAAAUGCUUCACUAAGUGAUCGAUUUGUGCUC__AUUUUGGAGUGCAGACUUUAC______UAAUAGAAUAUCUUCUUUUCAUUC .....................................((((((((........))))))))..................................... ( -7.18 = -6.48 + -0.70)


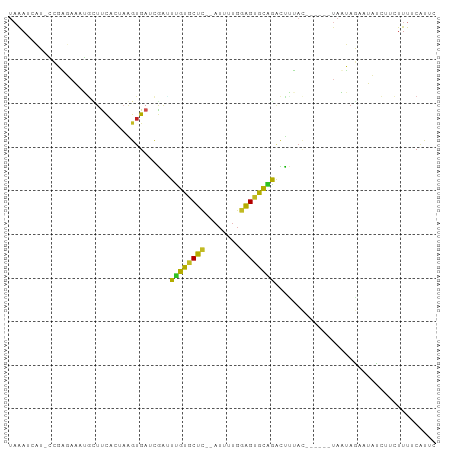
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:02:50 2011