
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 4,237,908 – 4,238,003 |
| Length | 95 |
| Max. P | 0.820293 |

| Location | 4,237,908 – 4,238,003 |
|---|---|
| Length | 95 |
| Sequences | 7 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 64.31 |
| Shannon entropy | 0.73065 |
| G+C content | 0.55098 |
| Mean single sequence MFE | -40.29 |
| Consensus MFE | -10.42 |
| Energy contribution | -10.77 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.55 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.820293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 4237908 95 + 24543557 GACCACCAGCUUGGUCAUUAUCGGUUCGGGAACCAC-AUGAUGAUG-----------GGACCUUGCCACCGCAGUUGGAAAUGGCUGUGAAAAUGGUCACCUAGCAA- (((((...((..((((((((((((((....))))..-..)))))).-----------.))))..))...(((((((......)))))))....))))).........- ( -29.90, z-score = -0.64, R) >dp4.chrXR_group8 8405911 104 + 9212921 GACCAGUACCUUUAUCAGCUCCAUCUUAAUGAUGAUGAUGAUGAUA--CUCGUAAUGAUGAG-AGCCGCU-CAUUGAUUAGCGUCUGUGGUUGGGGAGACCUUGCUAA ((((((((.(.((((((....((((.....)))).)))))).).))--))..((((((((((-.....))-)))).))))).)))..((((.(((....))).)))). ( -28.20, z-score = -1.14, R) >droEre2.scaffold_4784 6950235 94 + 25762168 GACCACCAGCUCGGUCAUUAUCUGGUAGGGAACCAC-AUGAUGAUGGCUUGGUGGCGGGACCUUGCCACCGCAGUUG------------GAAAUGGUCACCUAGCAA- ((((((((((((((((((((((((((.....)))).-..)))))))))).((((((((....))))))))).)))))------------)...))))).........- ( -41.80, z-score = -3.33, R) >droYak2.chr3L 4813724 106 + 24197627 GACCACCAGCUCGGUCAUUAUCUGGUAGGGAACCAC-AUGAUGACGGCCCGGUGGCGGGACCUUGCCACCGCAGCUGAAAAUGGCAGUGGAAAUGGUCACCUAGCAA- (((((((((((..((((((((.((((.....)))).-))))))))(((.(((((((((....)))))))))..)))......)))..)))...))))).........- ( -42.40, z-score = -1.97, R) >droSec1.super_2 4234553 106 + 7591821 GACCACCAGCUUGGUCAUUAUCGGUUCGGGAACCGC-AUGAUGAUGCCCUGGUGGCGGGACCUUGCCACCGCAGCUGGAAAUGGCUGUGGAAAUGGUCACCUAGCAA- (.(((((((....(((((((((((((....))))).-))))))))...))))))))((((((......((((((((......))))))))....)))).))......- ( -48.00, z-score = -3.15, R) >droSim1.chr3L 3777095 106 + 22553184 GACCACCAGCUUGGUCAUUAUCGGUUCGGGAACCAC-AUGAUGAUGCCCUGGUGGCGGGACCUUGCCACCGCAGCUGGAAAUGGCUGUGGAAAUGGUCACCUAGCAA- (.(((((((...((.(((((((((((....))))..-..)))))))))))))))))((((((......((((((((......))))))))....)))).))......- ( -45.10, z-score = -2.57, R) >droMoj3.scaffold_6500 13935251 105 - 32352404 GAGCUCCGGCUCGGGCUGUGCCCGUUCGACCACCGC-GCGCUGCCUGCUUUACAAUUGUGGGUGGC-ACCACAACUGGUGGUGUUGGAUUAGGUGGUGGCUGCACAG- ((((....))))((((...))))((.((.(((((((-..((..((..(.........)..))..))-(((((.....)))))..........))))))).)).))..- ( -46.60, z-score = -1.16, R) >consensus GACCACCAGCUUGGUCAUUAUCGGUUCGGGAACCAC_AUGAUGAUGGCCUGGUGGCGGGACCUUGCCACCGCAGCUGGAAAUGGCUGUGGAAAUGGUCACCUAGCAA_ (((((((((((..((((((((......(....)....))))))))...................((....))))))))...............))))).......... (-10.42 = -10.77 + 0.36)

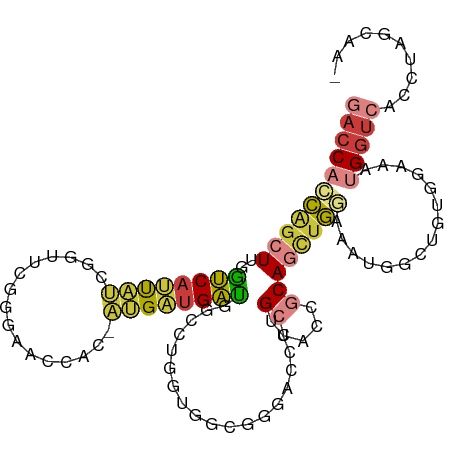

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:02:26 2011