
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 4,143,707 – 4,143,802 |
| Length | 95 |
| Max. P | 0.608964 |

| Location | 4,143,707 – 4,143,802 |
|---|---|
| Length | 95 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 62.10 |
| Shannon entropy | 0.79902 |
| G+C content | 0.55243 |
| Mean single sequence MFE | -25.92 |
| Consensus MFE | -6.84 |
| Energy contribution | -6.55 |
| Covariance contribution | -0.29 |
| Combinations/Pair | 1.88 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.608964 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
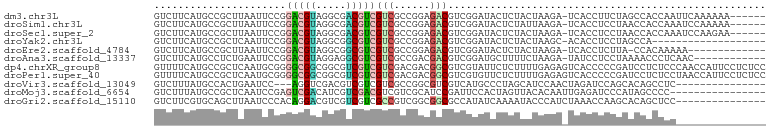
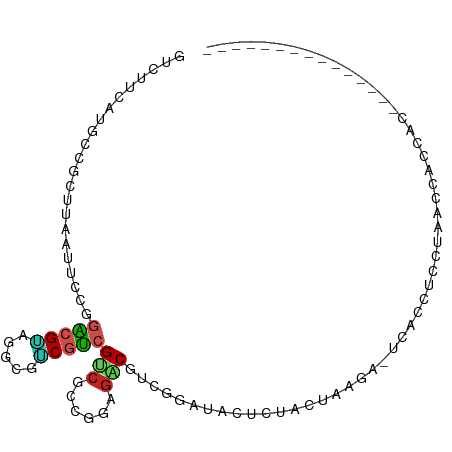
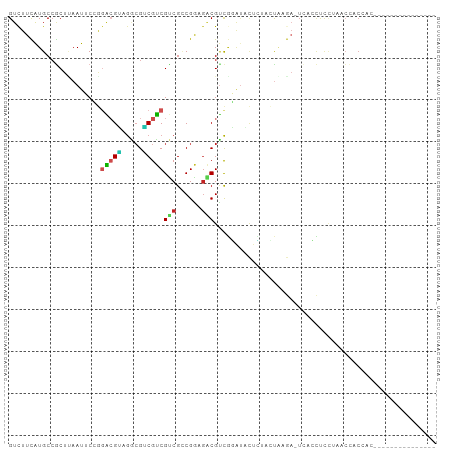
>dm3.chr3L 4143707 95 + 24543557 GUCUUCAUGCCGCUUAAUUCCGGACGUAGGCGACGUCGUCGCCGGAGACGUCGGAUACUCUACUAAGA-UCACCUUCUAGCCACCAAUUCAAAAAA------ ((((((((((((........))).))).((((((...))))))))))))...((((......(((.((-......)))))......))))......------ ( -24.50, z-score = -1.18, R) >droSim1.chr3L 3683725 95 + 22553184 GUCUUCAUGCCGCUUAAUUCCGGACGUAGGCGACGUCGUCGCCGGAGACGUCGGAUACUCUAUUAAGA-UCACCUCCUAACCACCAAAUCCAAAAA------ ((((((((((((........))).))).((((((...))))))))))))...((((......(((.((-.....)).))).......)))).....------ ( -24.52, z-score = -1.57, R) >droSec1.super_2 4137143 95 + 7591821 GUCUUCAUGCCGCUUAAUUCCGGACGUAGGCGACGUCGUCGCCGGAGACGUCGGAUACUCUACUAAGA-UCACCUCCUAACCACCAAAUCCAAGAA------ ((((((((((((........))).))).((((((...))))))))))))...((((.........((.-.......)).........)))).....------ ( -24.47, z-score = -1.00, R) >droYak2.chr3L 4714943 82 + 24197627 GUCUUCAUGCCGCUCAAUUCCGGACGUAGGCGGCGUCGUCGCCGGAGACGUCGGAUACUCUACUAAGC-ACACCUCCUAGCCA------------------- ((((((((((((........))).))).((((((...))))))))))))...(((.............-.....)))......------------------- ( -21.97, z-score = 0.44, R) >droEre2.scaffold_4784 6852577 87 + 25762168 GUCUUCAUGCCGCUUAAUUCCGGACGUAGGCGGCGUCGUCGCCGGAGACGUCGGAUACUCUACUAAGA-UCACCUCUUA-CCACAAAAA------------- ((((((((((((........))).))).((((((...))))))))))))((.((.........(((((-.....)))))-)))).....------------- ( -23.50, z-score = -0.85, R) >droAna3.scaffold_13337 14561188 89 + 23293914 GUCUUCAUGCCUCUGAAUUCCGGACGUAGGAGGCGUCGUCGCCGACGACGUCGGAUGCUUUUCUAAGA-UAUCCUCCUAAAACCCUCAAC------------ (((((((......))).....)))).((((((((((((((...)))))))))((((((((....))).-))))))))))...........------------ ( -28.60, z-score = -2.46, R) >dp4.chrXR_group8 8308051 102 + 9212921 GUUUUCAUGCCGCUCAAUGCGGGGCGGCGGCGUCGUCGUCGACGACGGCGUCGUAUUCUCUUUUGAGAGUCACCCCCGAUCCUCUCCCAACCAUUCCUCUCC ........(((((((......)))))))((((((((((....))))))))))((((((((....)))))).))............................. ( -38.00, z-score = -2.87, R) >droPer1.super_40 499347 102 + 810552 GUUUUCAUGCCGCUCAAUGCGGGGCGGCGGCGUCGUCGUCGACGACGGCGUCGUGUUCUCUUUUGAGAGUCACCCCCGAUCCUCUCCUAACCAUUCCUCUCC ........(((((((......)))))))((((((((((....))))))))))((((((((....))))).)))............................. ( -39.90, z-score = -3.19, R) >droVir3.scaffold_13049 1997688 84 + 25233164 GUCUUUAUGCCACUGAAUCC---AGGUCGACGUCGUCGUCGCCGGCGUCGUCAUGCCCUAGCAUCCAACUAGAUCCAGCACAGCCUC--------------- ........((..(((.(((.---((...((((.(((((....))))).))))((((....))))....)).))).)))....))...--------------- ( -21.80, z-score = -1.22, R) >droMoj3.scaffold_6654 727022 86 - 2564135 GUCUUUAUGCCGCUCAAUCCGAGUCGACAUCGUCGACGUCGUCGCAUCCGAUUCCACUAGUUACACAAUUGAGAUCCCAUAGCCCC---------------- ....(((((...((((((....((((((...))))))(..((((....))))..)............))))))....)))))....---------------- ( -17.80, z-score = -1.40, R) >droGri2.scaffold_15110 3058003 87 + 24565398 GUCUUCGUGCAGCUUAAUCCCACAGGACGUCGUCGUCGCCGUCGGCGGCGCCAUAUCAAAAUACCCAUCUAAACCAAGCACAGCUCC--------------- ......((((.......(((....)))((((((((.......))))))))...........................))))......--------------- ( -20.10, z-score = -1.42, R) >consensus GUCUUCAUGCCGCUUAAUUCCGGACGUAGGCGUCGUCGUCGCCGGAGACGUCGGAUACUCUACUAAGA_UCACCUCCUAACCACCAC_______________ ......................(((((.....)))))(((......)))..................................................... ( -6.84 = -6.55 + -0.29)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:02:13 2011