
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 3,440,527 – 3,440,608 |
| Length | 81 |
| Max. P | 0.727304 |

| Location | 3,440,527 – 3,440,608 |
|---|---|
| Length | 81 |
| Sequences | 10 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 76.24 |
| Shannon entropy | 0.44813 |
| G+C content | 0.50670 |
| Mean single sequence MFE | -20.45 |
| Consensus MFE | -11.29 |
| Energy contribution | -11.30 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.727304 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 3440527 81 - 24543557 --------------UGCCUGCCUGCACUUUAAAACC--AAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCACCGCCAAAUCCAAAUCCGUC --------------(((.((......((((......--))))......))))).(((((..(((((((((........))))...))))).))))). ( -18.90, z-score = -1.60, R) >droSim1.chr3L 2978091 81 - 22553184 --------------UGCCUGCCUGCACUUUAAAACC--AAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCACCGCCAAAUCCAAAUCCGUC --------------(((.((......((((......--))))......))))).(((((..(((((((((........))))...))))).))))). ( -18.90, z-score = -1.60, R) >droSec1.super_2 3432625 81 - 7591821 --------------UGCCUGCCUGCACUUUAAAACC--AAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCACCGCCAAAUCCAAAUCCGUC --------------(((.((......((((......--))))......))))).(((((..(((((((((........))))...))))).))))). ( -18.90, z-score = -1.60, R) >droYak2.chr3L 3988477 77 - 24197627 ------------------UGCCUGCAGUUUAAAACC--AAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCACCGCCAAAUCCAAAUCCGUC ------------------....(((.((((......--.......)))).))).(((((..(((((((((........))))...))))).))))). ( -19.12, z-score = -2.21, R) >droEre2.scaffold_4784 6130493 81 - 25762168 --------------UGCCUGCCUGCACUUUAAAACC--AAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCACCGCCAAAUCCAAAUCCGUC --------------(((.((......((((......--))))......))))).(((((..(((((((((........))))...))))).))))). ( -18.90, z-score = -1.60, R) >droAna3.scaffold_13337 13294463 82 + 23293914 ---------------UGCGGCCCGCACUUAAAAACCAAAAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCAGCGCCAAAUCCAAAUCCGUC ---------------((((...))))...............((.......))..(((((..(((((((((((....))))))...))))).))))). ( -21.60, z-score = -1.87, R) >dp4.chrXR_group8 5547917 93 - 9212921 UCCCCGCGGUUUCUGAGGGGCCUGUGCACUUAAA----AACAGCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCAGCGCCAAAUCCAAAUCCGUC .((((.(((...))).)))).((((.........----.))))...........(((((..(((((((((((....))))))...))))).))))). ( -29.30, z-score = -1.41, R) >droWil1.scaffold_180955 1191178 73 - 2875958 ----------------UGUGUGUAUAUUCUUUACAC--AAAACCAAAACAGCAAACGGAAGUUGGAGGCGUUUCCCAGCGCCAAAUCCGUC------ ----------------..((((((.......)))))--)..............(((....)))((((((((......)))))...)))...------ ( -19.30, z-score = -2.25, R) >droGri2.scaffold_15110 5850227 68 - 24565398 --------------------CUUGCAUUUUAAAAGC--CGUAGC-AAACAGCCAACGGAAGUUGGAGGCGUUACUCAGAGCCAAAUCUGGC------ --------------------..............((--((((((-...(..(((((....)))))..).)))))..(((......))))))------ ( -18.90, z-score = -1.30, R) >droVir3.scaffold_13049 18267079 73 + 25233164 --------------UGCCUGCCUGCA-UUUAAAGGC--CGAACC-AAACAGCGAACGGAAGUUGGAGGCGUUGCUCAGCGCCAAAUCCGAC------ --------------.....((((...-.....))))--((....-......(.(((....))).).((((((....)))))).....))..------ ( -20.70, z-score = -0.30, R) >consensus ______________UGCCUGCCUGCACUUUAAAACC__AAAGCCAAAACAGCAAACGGAAGUUGGAGGCGUUACCCACCGCCAAAUCCAAAUCCGUC .....................................................(((....)))(((((((........))))...)))......... (-11.29 = -11.30 + 0.01)


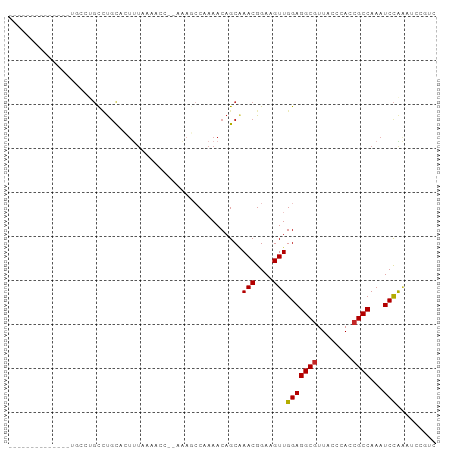
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:00:44 2011