
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 3,399,244 – 3,399,350 |
| Length | 106 |
| Max. P | 0.784393 |

| Location | 3,399,244 – 3,399,350 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 79.74 |
| Shannon entropy | 0.35253 |
| G+C content | 0.35884 |
| Mean single sequence MFE | -18.55 |
| Consensus MFE | -13.28 |
| Energy contribution | -13.72 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.784393 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
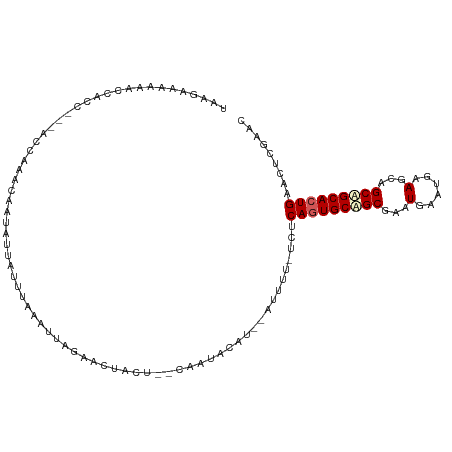
>dm3.chr3L 3399244 106 - 24543557 AAAGAAAAAUCCACCACCACCAAAGAAUAUUAUUUAAAUUAGAACUACU--UGAUACAU--AUUUU-UCUCAGUGCAGCGAAUGAAUGAAGCAGCAGCACUGAACUCGAAC ...((.................((((((((((((...............--.)))).))--)))))-).(((((((.((...(......)...)).)))))))..)).... ( -14.29, z-score = -1.03, R) >droSim1.chr3L 2934943 102 - 22553184 UUAGAACAAAUUAAC---ACCAAACAAUAUUAUUUAAAUUUGAACUGCU--CAAUACAU---UUUU-UCUCAGUGCAGCGAAUGAAUGAAGCAGCAGCACUGAACUCGAAC .(((..((((((...---..................))))))..)))..--........---....-..(((((((.((...(......)...)).)))))))........ ( -13.40, z-score = -0.03, R) >droSec1.super_2 3392068 105 - 7591821 UAAGAAAAAACCACCUUAACAAAACAAUAUUAUUAAAAUUUGAACUGCU--CAAUACAU---UUUU-UCUCAGUGCUGCGAAUGAAUGAAGCAGCAGCACUGAACUCGAAC ...(((((((.................((....))....((((.....)--)))....)---))))-))((((((((((...(......)...))))))))))........ ( -19.80, z-score = -2.71, R) >droYak2.chr3L 3946549 103 - 24197627 UAAGAAGAAAACACC---ACCUAACUA---UAUUUAAAGUAGAACUACUAGUUAUACAU--AUUUUCUGCCAGUGCAGCGAAUGAAUGAAGCAGCUGCACUGAACUCGAAC ...((((((((....---...(((((.---.......((((....))))))))).....--.))))))..(((((((((...(......)...)))))))))...)).... ( -21.52, z-score = -2.56, R) >droEre2.scaffold_4784 6089533 105 - 25762168 UAUGUGAGAGCCACC---ACCAAAAAA---UAUUUGAGUUAGAACUACUAGAUAAACAUGCAUUUUCUGGCAGUGCAGCGAAUGAAUGAAGCAGCUGCAGUGAACUCGAAC ...((....))....---.........---..((((((((.......(((((.............)))))((.((((((...(......)...)))))).)))))))))). ( -23.72, z-score = -1.30, R) >consensus UAAGAAAAAACCACC___ACCAAACAAUAUUAUUUAAAUUAGAACUACU__CAAUACAU__AUUUU_UCUCAGUGCAGCGAAUGAAUGAAGCAGCAGCACUGAACUCGAAC ......................................................................(((((((((...(......)...)))))))))......... (-13.28 = -13.72 + 0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:00:34 2011