
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 3,228,939 – 3,229,042 |
| Length | 103 |
| Max. P | 0.575290 |

| Location | 3,228,939 – 3,229,042 |
|---|---|
| Length | 103 |
| Sequences | 8 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 77.60 |
| Shannon entropy | 0.45361 |
| G+C content | 0.56585 |
| Mean single sequence MFE | -37.59 |
| Consensus MFE | -20.18 |
| Energy contribution | -21.26 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.575290 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 3228939 103 + 24543557 UCAGCCUCCCAUAUCCAGGGCGGUAACAUCAUGCCCAUCCAGAUAACUGCACCCAUUCCAGGAGGAAUGGUUAGCGUGGUUCCGACCAACAUUGUGCAGGAUC--- ...(((.(((.......))).)))......................(((((((((((((....)))))))......((((....)))).....))))))....--- ( -31.30, z-score = -1.03, R) >droSim1.chr3L 2761570 103 + 22553184 UCAGCCUCCCACAUCCAGGGCGGUAACAUCGUGCCCAUCCAGAUAACUGGACCCAUUCCAGGAGGAAUGGUCAGUGUGGUCCCGACCAACAUUGUGCAGGAUC--- ...(((.(((.......))).)))...(((.(((...(((((....))))).(((((((....))))))).(((((((((....))).)))))).))).))).--- ( -38.30, z-score = -1.95, R) >droSec1.super_2 3222866 103 + 7591821 UCAGCCUCCCACAUCCAGGGCGGUAACAUCGUGCCCAUCCAGAUAACUGGACCCAUUCCAGGAGGAAUGGUCAGUGUGGUCCCGACCAACAUUGUGCAGGAUC--- ...(((.(((.......))).)))...(((.(((...(((((....))))).(((((((....))))))).(((((((((....))).)))))).))).))).--- ( -38.30, z-score = -1.95, R) >droYak2.chr3L 3773527 103 + 24197627 ACAGCCUCGCACAUCCAGGGCGGCAGUAUUGUGCCCAUCCAGAUAACUGGUGCCAUUCCAGGAGGAAUGGUCAGCGUGGUCCCGGCCAACAUUGUGCAGGAUC--- ....(((.(((((....((.(((....((..(((....((((....)))).((((((((....))))))))..)))..)).))).)).....))))))))...--- ( -39.10, z-score = -1.26, R) >droEre2.scaffold_4784 5911629 103 + 25762168 ACAGCCUCGCACAUCCAGGGCGGUAAUAUUGUACCCAUCCAGAUAACUGGAGCUAUUCCAGGAGGAAUGGUCAGCGUGGUCCCGGCCAACAUUGUGCAGGAUC--- ....(((.(((((.....(((.(..((..(((.....(((((....)))))((((((((....))))))))..)))..))..).))).....))))))))...--- ( -35.00, z-score = -1.19, R) >droAna3.scaffold_13337 7533171 106 + 23293914 AAUCCACCGCACCUCCAGGCCGGCAGCAUCGUGCCGAUCCAGAUAACUGGCGCCAUUCCGGGUGGAAUGGUGAGCGUACUGCCUGCAAACAUAGUGCAGGGUGAUC ....((((((((......((.(((((...(((......((((....))))(((((((((....))))))))).)))..))))).)).......))))..))))... ( -42.12, z-score = -1.54, R) >dp4.chrXR_group8 5890104 100 - 9212921 ACUCCAUCCCAUAUCCAGAGCGGCAGCAUUGUGCCCAUCCAGCUAGCGGGAGGCAUUCCCGGCGGCAUGGUGAGCGUGGUGCCCGCCAAUAUUGUGGAUC------ ........(((((......((((..((((..(((.((.((((((..((((((...))))))..))).))))).)))..))))))))......)))))...------ ( -40.40, z-score = -1.11, R) >droWil1.scaffold_180698 8812024 97 + 11422946 GCUGGUGCUCAUAUCCAGGGUGGCAGUAUUGUGCCAAUACAGCUAUCAGGAG---UACCGGGUGGUAUGGUGAGCGUGGUGCCAGCCACAAUUGUAGAUC------ ((((((((((((.(((..((((((.((((((...)))))).)))))).)))(---((((....))))).)))))).....))))))..............------ ( -36.20, z-score = -1.27, R) >consensus ACAGCCUCCCACAUCCAGGGCGGCAACAUCGUGCCCAUCCAGAUAACUGGAGCCAUUCCAGGAGGAAUGGUCAGCGUGGUCCCGGCCAACAUUGUGCAGGAUC___ ....(((.(((((......((((((......))))..(((((....))))).(((((((....)))))))...)).((((....))))....))))))))...... (-20.18 = -21.26 + 1.08)

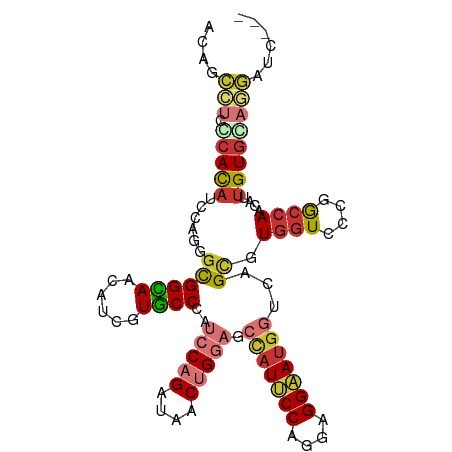
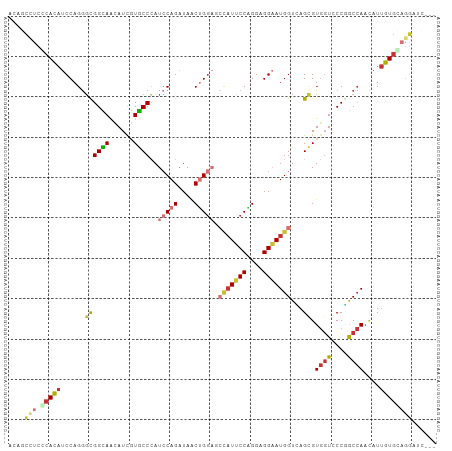
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:00:08 2011