
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 3,141,726 – 3,141,830 |
| Length | 104 |
| Max. P | 0.656545 |

| Location | 3,141,726 – 3,141,830 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 94.92 |
| Shannon entropy | 0.08893 |
| G+C content | 0.47412 |
| Mean single sequence MFE | -30.50 |
| Consensus MFE | -28.40 |
| Energy contribution | -28.64 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.93 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.656545 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
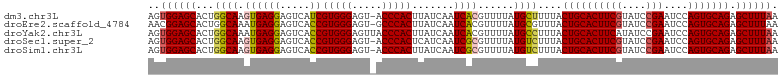
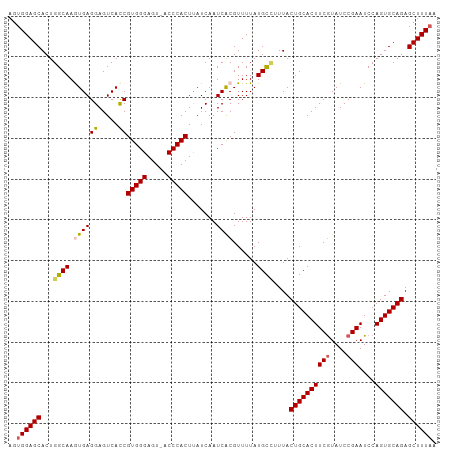
>dm3.chr3L 3141726 104 + 24543557 AGUGGAGCACUGGCAAGUGAGGAGUCAUCGUGGGAGU-ACCCACUUAUCAAUCACGUUUUAUGCUUUUACUGCACUUCGUAUCCGAAUCCAGUGCAGAGCUUUAA ..((((((...((((.((((.((....))(((((...-.))))).......))))......))))....((((((((((....)))....))))))).)))))). ( -30.60, z-score = -1.47, R) >droEre2.scaffold_4784 3150777 104 + 25762168 AACGGAGCACUGGCAAAUGAGGAGUCACCGUGGGAGU-GCCCACUUAUCAAUCACGUUUUAUGCGUUUACUGCACUUCGUAUCCGAAUCCAGUGCAGAGCUUUAA ...(((((....(((((((.((.....))(((((...-.)))))..........))))...))).....((((((((((....)))....))))))).))))).. ( -28.80, z-score = -0.77, R) >droYak2.chr3L 15396531 105 - 24197627 AGUGGAGCACUGGCAAAUGAGGAGUCACCGUGGGAGUUACCCACUUAUCAAUCACGUUUUAUGCCUUUACUGCACUUCAUAUCCGAAUCCAGUGCAGAGCUUUAA ..((((((...((((((((.((.....))(((((.....)))))..........))))...))))....(((((((((......))....))))))).)))))). ( -30.70, z-score = -1.86, R) >droSec1.super_2 3135348 104 + 7591821 AGUGGAGCACUGGCAAGUGAGGAGUCACCGUGGGAGU-ACCCACUCAUCAAUCGCGUUUUAUGUCUUUACUGCACUUCGUAUCCGAAUCCAGUGCAGAGCUUUAA ..((((((...((((.((((((.....))(((((...-.))))).......))))......))))....((((((((((....)))....))))))).)))))). ( -31.20, z-score = -1.20, R) >droSim1.chr3L 2669323 104 + 22553184 AGUGGAGCACUGGCAAGUGAGGAGUCACCGUGGGAGU-ACCCACUUAUCAAUCGCGUUUUAUGUCUUUACUGCACUUCGUAUCCGAAUCCAGUGCAGAGCUUUAA ..((((((...((((.((((((.....))(((((...-.))))).......))))......))))....((((((((((....)))....))))))).)))))). ( -31.20, z-score = -1.34, R) >consensus AGUGGAGCACUGGCAAGUGAGGAGUCACCGUGGGAGU_ACCCACUUAUCAAUCACGUUUUAUGCCUUUACUGCACUUCGUAUCCGAAUCCAGUGCAGAGCUUUAA ..((((((...((((.((((((.....))(((((.....))))).......))))......))))....((((((((((....)))....))))))).)))))). (-28.40 = -28.64 + 0.24)

| Location | 3,141,726 – 3,141,830 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 94.92 |
| Shannon entropy | 0.08893 |
| G+C content | 0.47412 |
| Mean single sequence MFE | -29.76 |
| Consensus MFE | -26.82 |
| Energy contribution | -26.42 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.607761 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
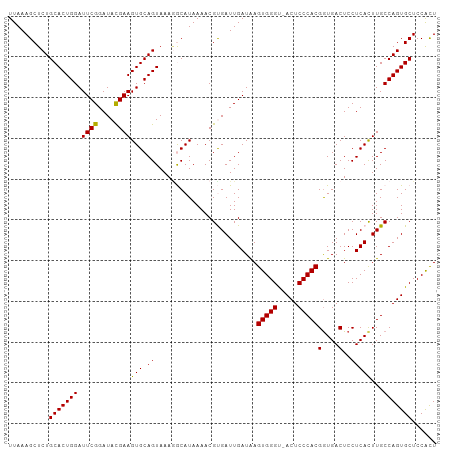
>dm3.chr3L 3141726 104 - 24543557 UUAAAGCUCUGCACUGGAUUCGGAUACGAAGUGCAGUAAAAGCAUAAAACGUGAUUGAUAAGUGGGU-ACUCCCACGAUGACUCCUCACUUGCCAGUGCUCCACU ..........(((((((.((((....))))((((.......)))).....((((.......(((((.-...))))).........))))...)))))))...... ( -30.19, z-score = -2.21, R) >droEre2.scaffold_4784 3150777 104 - 25762168 UUAAAGCUCUGCACUGGAUUCGGAUACGAAGUGCAGUAAACGCAUAAAACGUGAUUGAUAAGUGGGC-ACUCCCACGGUGACUCCUCAUUUGCCAGUGCUCCGUU ....(((...(((((((.((((....))))((((.......)))).....((((.......(((((.-...)))))(....)...))))...)))))))...))) ( -29.60, z-score = -1.11, R) >droYak2.chr3L 15396531 105 + 24197627 UUAAAGCUCUGCACUGGAUUCGGAUAUGAAGUGCAGUAAAGGCAUAAAACGUGAUUGAUAAGUGGGUAACUCCCACGGUGACUCCUCAUUUGCCAGUGCUCCACU ....(((.(((((((..((......))..)))))))....((((......((((.......(((((.....)))))(....)...)))).))))...)))..... ( -28.90, z-score = -0.99, R) >droSec1.super_2 3135348 104 - 7591821 UUAAAGCUCUGCACUGGAUUCGGAUACGAAGUGCAGUAAAGACAUAAAACGCGAUUGAUGAGUGGGU-ACUCCCACGGUGACUCCUCACUUGCCAGUGCUCCACU ....(((.(((.((((.(((((....)).))).)))).............((((.(((.(((((((.-...)))))(....))).))).))))))).)))..... ( -30.80, z-score = -1.59, R) >droSim1.chr3L 2669323 104 - 22553184 UUAAAGCUCUGCACUGGAUUCGGAUACGAAGUGCAGUAAAGACAUAAAACGCGAUUGAUAAGUGGGU-ACUCCCACGGUGACUCCUCACUUGCCAGUGCUCCACU ....(((.(((.((((.(((((....)).))).)))).............((((.(((...(((((.-...)))))(....)...))).))))))).)))..... ( -29.30, z-score = -1.41, R) >consensus UUAAAGCUCUGCACUGGAUUCGGAUACGAAGUGCAGUAAAGGCAUAAAACGUGAUUGAUAAGUGGGU_ACUCCCACGGUGACUCCUCACUUGCCAGUGCUCCACU ..........(((((((.((((....))))(((.((.........................(((((.....)))))(....)..)))))...)))))))...... (-26.82 = -26.42 + -0.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:59:55 2011