
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 3,083,198 – 3,083,342 |
| Length | 144 |
| Max. P | 0.603806 |

| Location | 3,083,198 – 3,083,342 |
|---|---|
| Length | 144 |
| Sequences | 6 |
| Columns | 152 |
| Reading direction | reverse |
| Mean pairwise identity | 70.37 |
| Shannon entropy | 0.56508 |
| G+C content | 0.38219 |
| Mean single sequence MFE | -35.80 |
| Consensus MFE | -13.52 |
| Energy contribution | -13.22 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.603806 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 3083198 144 - 24543557 UGAGCCGCCAAACUUAGAACUUGGCAAUUUGCACAUUUUAAAGGAAAUAUAAUAAUACAACAGCAUACAUAUUUGUA-UGUA----GUAUAAUCGGUAUAUGC---GUGCGUGCUACGGUCAGGCUGCUCAGGUGUUAAUCACUAUACCUUC (((((.(((..(((.......(((((...(((((............................(((((((....))))-))).----(((((.......)))))---)))))))))).)))..))).)))))(((((........)))))... ( -37.50, z-score = -1.48, R) >droWil1.scaffold_181017 153053 148 - 939937 GAAGAAAACAAA-UUAACAUUUUUCUACAUAUUCCUUCAUUUAGAAUUCCCUUCCUACUAGCGUAAACAUAAUGGUUAGAUCUUUUCCUGAUCUUGCCAAUGU---UCACUCGUUACUUGCCAACUCGCUACUCGUUAAUUGCUAUAAAUAC ..........((-(((((....(((((.((........)).)))))............((((((.(((((..((((.(((((.......))))).))))))))---).))..(((.......)))..))))...)))))))........... ( -24.40, z-score = -2.92, R) >droSim1.chr3L 2606097 137 - 22553184 UGAGCCGCCAAACUUAGAACUUGGCAAUUUGCACAUUUUAAAGAAAAUAUAAUA----------AUACAUAUUUGUA-UGUA----GUAUAUUCGGUAUGUGCGUCGUGCGUGCUGCGGUCAGGCUGCUCAGGUGUUAAUCACUAUACCUUC (((((.(((..((((((.((...(((...(((((((......(((.((((....----------(((((....))))-)...----)))).)))...)))))))...))))).))).)))..))).)))))(((((........)))))... ( -39.50, z-score = -2.09, R) >droSec1.super_2 3076732 147 - 7591821 UGAGCCGCCAAACUUAGAACUUGGCAAUUUGCACAUUUUAAAGAAAAUAUAAUAAUACAACAGCAUACAUAUUUGUA-UGUA----GUAUAUUCGGUAUGUGCGUCGUGCGUGCUGCGGUCAGGCUGCUCAGGUGUUAAUCACUAUACCUUC (((((.(((..((((((.((...(((...(((((((......(((..(((....)))..((.(((((((....))))-))).----))...)))...)))))))...))))).))).)))..))).)))))(((((........)))))... ( -40.20, z-score = -1.49, R) >droYak2.chr3L 15333312 148 + 24197627 UGAGCCGCCAAACUUAGAACUUGGCAAUUUGCACAUUUUAAAGGAAAUAUAAUAAUACAACCGCAUAUAUUAUAGUAGCAUGUAGUAUGUAGUCUGUAUGUGC---GUG-UUGUUACGGUCAGGUUGCUCAGGUGUUAAUCACUAUACCUUC (((((.(((..(((.......(((((((.((((((((((....)))))......(((((..(..((((((((((......)))))))))).)..)))))))))---).)-)))))).)))..))).)))))(((((........)))))... ( -39.40, z-score = -1.54, R) >droEre2.scaffold_4784 3091406 138 - 25762168 GGAGCCGCCAAACUUAGAACUUGGCAAUUUGCACAUUUUAAAGGAAAUAUAAUAAAACAUCAGCCUACAGUAGUGUAGUG------GUAUAGGC-GUAUGUGC---G----UGUUUCGGUCAGGUUGCUCAGGUGUUAAUCACUAUACCUUC .((((.(((...(((.(((..((((.....)).)).))).)))(((((((......((((..(((((.(.((......))------.).)))))-..))))..---)----)))))).....))).))))((((((........)))))).. ( -33.80, z-score = -0.37, R) >consensus UGAGCCGCCAAACUUAGAACUUGGCAAUUUGCACAUUUUAAAGGAAAUAUAAUAAUACAACAGCAUACAUAAUUGUA_UGUA____GUAUAUUCGGUAUGUGC___GUGCGUGCUACGGUCAGGCUGCUCAGGUGUUAAUCACUAUACCUUC ..(((((((((.........))))).....((((((((.....)))...................(((......))).....................)))))...................))))....((((((........)))))).. (-13.52 = -13.22 + -0.30)

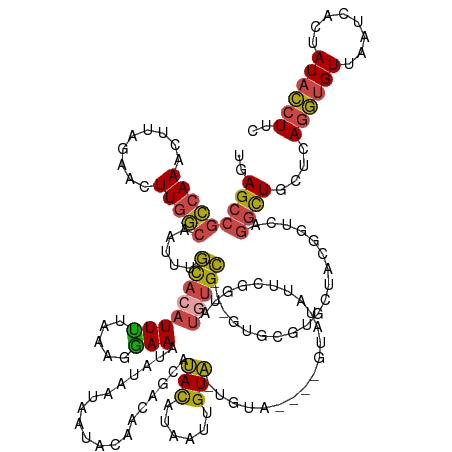
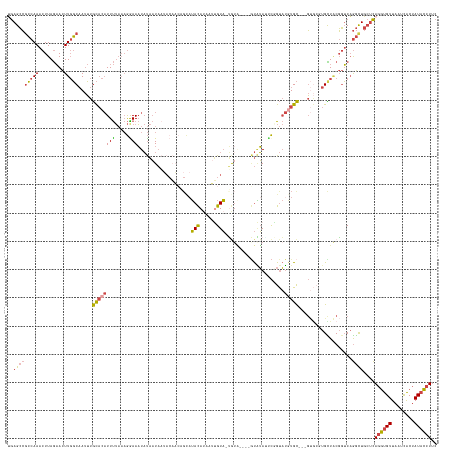
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:59:37 2011