
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 2,920,139 – 2,920,196 |
| Length | 57 |
| Max. P | 0.628732 |

| Location | 2,920,139 – 2,920,196 |
|---|---|
| Length | 57 |
| Sequences | 12 |
| Columns | 63 |
| Reading direction | reverse |
| Mean pairwise identity | 78.42 |
| Shannon entropy | 0.44482 |
| G+C content | 0.51590 |
| Mean single sequence MFE | -17.68 |
| Consensus MFE | -10.38 |
| Energy contribution | -10.39 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.628732 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 2920139 57 - 24543557 CUUUUCGGGGCU--UUC-UUUUGCGUGCC-ACGGCGACUUAUUGGCCAUUGGCAUGCAAGC-- ............--...-.((((((((((-(.(((.(.....).)))..))))))))))).-- ( -20.20, z-score = -2.13, R) >droSim1.chr3L 2473184 57 - 22553184 CUUUUCGGGGCU--UUC-UUUUGCGUGCC-ACGGCGACUUAUUGGCCAUUGGCAUGCAAGC-- ............--...-.((((((((((-(.(((.(.....).)))..))))))))))).-- ( -20.20, z-score = -2.13, R) >droSec1.super_2 2939889 57 - 7591821 CUUUUCGGGGCU--UUC-UUUUGCGUGCC-ACGGCGACUUAUUGGCCAUUGGCAUGCAAGC-- ............--...-.((((((((((-(.(((.(.....).)))..))))))))))).-- ( -20.20, z-score = -2.13, R) >droYak2.chr3L 15189809 57 + 24197627 CUUUUCGGGGCU--UUC-UUUUGCGUGCC-ACGGCGACUUAUUGGCCACUGGCAUGCAAGC-- ............--...-.((((((((((-(.(((.(.....).)))..))))))))))).-- ( -20.20, z-score = -2.10, R) >droEre2.scaffold_4784 2950129 57 - 25762168 CUUUUUGGGGCU--UUC-UUUUGCGUGCC-ACGGCGACUUAUUGGCCAUUGGCAUGCAAGC-- ............--...-.((((((((((-(.(((.(.....).)))..))))))))))).-- ( -20.20, z-score = -2.15, R) >droAna3.scaffold_13337 14122363 57 + 23293914 -UUUUCGGGGCU--UUCAUUUUGCGUGCC-ACAGCGACUUAUUGGCCAUUGGCAUGCAAGC-- -...........--.....((((((((((-(..((.(.....).))...))))))))))).-- ( -16.90, z-score = -1.07, R) >dp4.chrXR_group8 3757167 63 - 9212921 GUUUUUGGCUGUGGUCCGUUUUGCGUGCCAAAUGCGACUUAUUGGCCAUUGGCAUGCAAGGGC .............((((...(((((((((((..((.(.....).))..))))))))))))))) ( -20.90, z-score = -0.78, R) >droPer1.super_1450 1293 63 + 6820 GUUUUUGGCUGUGGUCCGUUUUGCGUGCCAAAUGCGACUUAUUGGCCAUUGGCAUGCAAGGGC .............((((...(((((((((((..((.(.....).))..))))))))))))))) ( -20.90, z-score = -0.78, R) >droWil1.scaffold_180698 390361 53 + 11422946 -CUUUUUGGGUU------UUUUGCGUGCG-AAUGCGAAUUAUUAGCCAUUGGCAUGCUAAC-- -...........------....((((((.-(((((.........).)))).))))))....-- ( -13.40, z-score = -1.44, R) >droGri2.scaffold_15110 19489774 50 - 24565398 --------GUUU--UGGUUUUUGCGUGCG-AACUCGACUUAUUGGCCAUUGGCAUGCAAGC-- --------....--.....(((((((((.-((.((((....))))...)).))))))))).-- ( -12.00, z-score = -0.39, R) >droMoj3.scaffold_6680 2952285 50 + 24764193 --------GUUU--UGGUUUUUGCGUGCG-AGCCCGACUUAUUGGCCAUUGGCAUGCAAGC-- --------....--.....(((((((((.-((.((((....))))...)).))))))))).-- ( -14.20, z-score = -0.54, R) >droVir3.scaffold_13049 15074512 50 - 25233164 --------GUUU--UGGUUUUUGCGUGCG-AGCUCGACUUAUUGGCCAUUGGCAUGCAAGC-- --------....--.....(((((((((.-(((.(((....)))))...).))))))))).-- ( -12.90, z-score = -0.21, R) >consensus _UUUUCGGGGCU__UUC_UUUUGCGUGCC_ACGGCGACUUAUUGGCCAUUGGCAUGCAAGC__ ...................(((((((((.......................)))))))))... (-10.38 = -10.39 + 0.01)


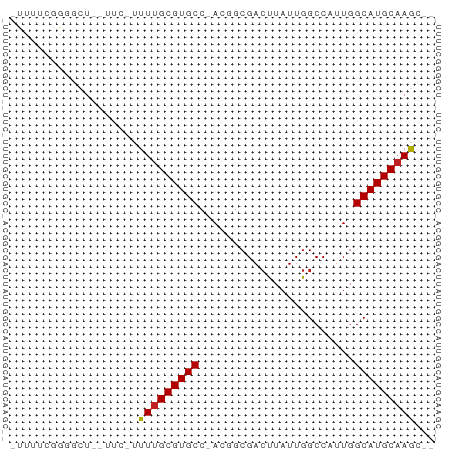
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:59:04 2011