
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 2,700,194 – 2,700,344 |
| Length | 150 |
| Max. P | 0.951680 |

| Location | 2,700,194 – 2,700,314 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.33 |
| Shannon entropy | 0.27328 |
| G+C content | 0.37567 |
| Mean single sequence MFE | -29.30 |
| Consensus MFE | -20.16 |
| Energy contribution | -19.88 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.751170 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 2700194 120 - 24543557 UUACAGUAACAUUUGAAAAAUGUCCUUGCUAAUUGGCAGCAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACGAAGCAGAUAUGUUUCUAAAUUCAUAAAUUGUACCAUACAAUU .((((((......((((...((.(.(((((...((((((((((.(((.....))).))))))))))))))).).)).((((((....))))))....))))...)))))).......... ( -33.80, z-score = -3.51, R) >droEre2.scaffold_4784 2724645 119 - 25762168 -UACAACAAGAUUUGGUUGAUUUCCUUGCAAAUUGGCAGCACAUGUUUGCUCGACCAGUGCUGCCAAGCUCAGCUACAAAGCAGAAUUGCUGCCUAAUUCAUAAAUUGUACCAUAAAAUC -(((((.......(((((((.......((...((((((((((.((.((....)).)))))))))))))))))))))....((((.....))))............))))).......... ( -29.81, z-score = -1.32, R) >droYak2.chr3L 14965016 120 + 24197627 CUUUAGCAAGAUUUGAUUGAUUUCCUUGCAAAUUGGCAUCACUUGUUUGCCCGACCAGUGCUGCCAAGCUAGGCUACAAAGCAGAUCUGUUUCCUAACUCAAAAUUUUAACCAUAAAAUU ....((((.((((((........(((.((...((((((.((((.(((.....))).)))).)))))))).)))........))))))))))...........(((((((....))))))) ( -27.39, z-score = -1.65, R) >droSec1.super_2 2726005 120 - 7591821 CUACAGUAACAUUUGAAAAAUGUCCUUGCUAAUUGGCAGGAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACAAAGCAGAUCUGUUUCUAAAUUCAUAAAUUGUACCAUACAAUU ..((((....(((((.....((.(.(((((...((((((.(((.(((.....))).))).))))))))))).).)).....))))))))).............(((((((...))))))) ( -26.10, z-score = -0.95, R) >droSim1.chr3L 2256334 120 - 22553184 CUACAGUAACAUUUGAAUAAAGUCCUUGCUAAUUGGCAGCAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACAAAGCAGAUCUGAUUCUAAAUUCAUAAAUUGUACCAUACAAUU .((((((......(((((...(.(.(((((...((((((((((.(((.....))).))))))))))))))).).).......(((......)))..)))))...)))))).......... ( -29.40, z-score = -2.34, R) >consensus CUACAGUAACAUUUGAAUAAUGUCCUUGCUAAUUGGCAGCAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACAAAGCAGAUCUGUUUCUAAAUUCAUAAAUUGUACCAUACAAUU .((((((.(((((.....)))))..((((...(((((((((((.(((.....))).)))))))))))((...))......))))....................)))))).......... (-20.16 = -19.88 + -0.28)

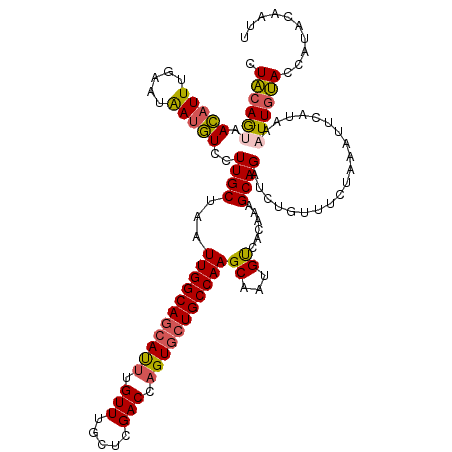
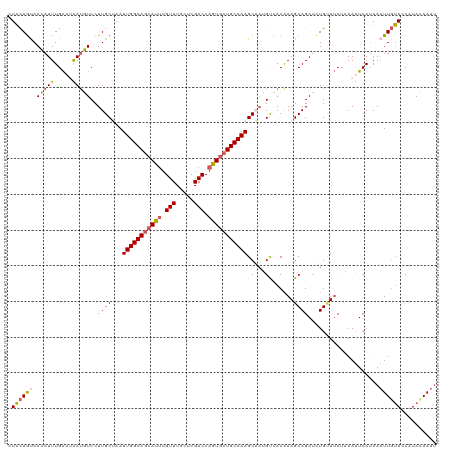
| Location | 2,700,224 – 2,700,344 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 70.75 |
| Shannon entropy | 0.51725 |
| G+C content | 0.41406 |
| Mean single sequence MFE | -29.86 |
| Consensus MFE | -18.00 |
| Energy contribution | -19.84 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.908913 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 2700224 120 + 24543557 ACAUAUCUGCUUCGUGACAUUGCUUGGCAGCACUGGUCGAGCAAACAAAUGCUGCCAAUUAGCAAGGACAUUUUUCAAAUGUUACUGUAAAAGAAAGAAAAUUUCUGUACAACAGAAGAC .........(((((((((((((((((((((((...((.......))...))))))))...)))..(((.....))).))))))))((((..(((((.....))))).))))...)))).. ( -32.50, z-score = -2.44, R) >droEre2.scaffold_4784 2724675 91 + 25762168 GCAAUUCUGCUUUGUAGCUGAGCUUGGCAGCACUGGUCGAGCAAACAUGUGCUGCCAAUUUGCAAGGAAAUCAACCAAAUCUUGUUGUAGC----------------------------- (((((((.(((....))).))).((((((((((((..........)).)))))))))).))))........((((........))))....----------------------------- ( -26.60, z-score = -0.36, R) >droYak2.chr3L 14965046 111 - 24197627 ACAGAUCUGCUUUGUAGCCUAGCUUGGCAGCACUGGUCGGGCAAACAAGUGAUGCCAAUUUGCAAGGAAAUCAAUCAAAUCUUGCUAAAGGACAUAAACACAAGAGUGGGC--------- .....((..((((...(((..(((.(......).)))..)))......((((((((.....((((((............))))))....)).)))...))).))))..)).--------- ( -25.80, z-score = 0.64, R) >droSec1.super_2 2726035 120 + 7591821 ACAGAUCUGCUUUGUGACAUUGCUUGGCAGCACUGGUCGAGCAAACAAAUCCUGCCAAUUAGCAAGGACAUUUUUCAAAUGUUACUGUAGAAGAAAGAACUUUUCUGUACAGCAGAAAAC .....(((((...(((((((((((((((.......))))))).......((((((......)).)))).........)))))))).))))).........(((((((.....))))))). ( -34.80, z-score = -2.41, R) >droSim1.chr3L 2256364 104 + 22553184 UCAGAUCUGCUUUGUGACAUUGCUUGGCAGCACUGGUCGAGCAAACAAAUGCUGCCAAUUAGCAAGGACUUUAUUCAAAUGUUACUGUAGAAUAAAGAAAUUUU---------------- .....(((((...(((((((((((((((((((...((.......))...))))))))...)))..(((.....))).)))))))).))))).............---------------- ( -29.60, z-score = -2.31, R) >consensus ACAGAUCUGCUUUGUGACAUUGCUUGGCAGCACUGGUCGAGCAAACAAAUGCUGCCAAUUAGCAAGGACAUUAUUCAAAUGUUACUGUAGAAGAAAGAAAUUUU________________ .....(((((...(((((((((((((((((((...((.......))...)))))))))...))..(((.....))).)))))))).)))))............................. (-18.00 = -19.84 + 1.84)

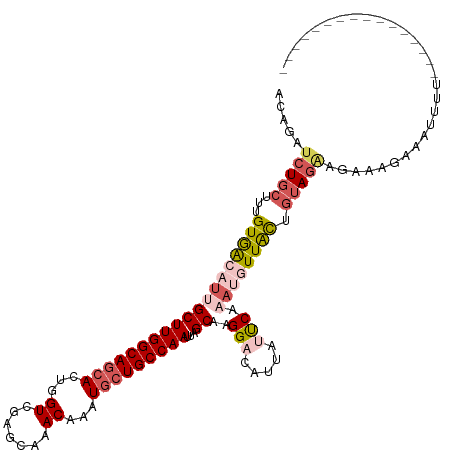
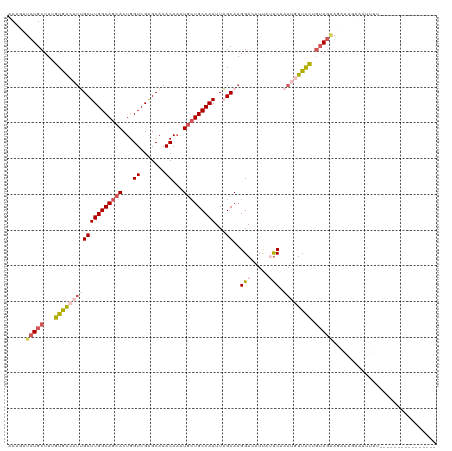
| Location | 2,700,224 – 2,700,344 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 70.75 |
| Shannon entropy | 0.51725 |
| G+C content | 0.41406 |
| Mean single sequence MFE | -28.86 |
| Consensus MFE | -19.23 |
| Energy contribution | -20.19 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.951680 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
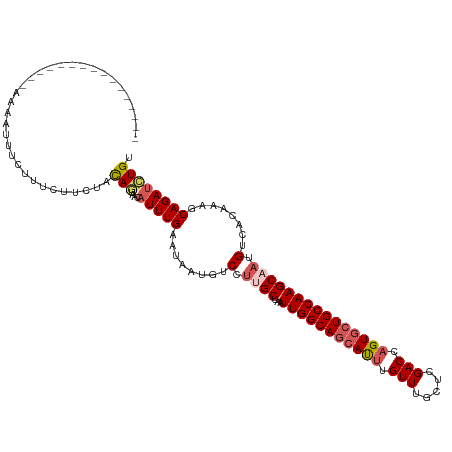
>dm3.chr3L 2700224 120 - 24543557 GUCUUCUGUUGUACAGAAAUUUUCUUUCUUUUACAGUAACAUUUGAAAAAUGUCCUUGCUAAUUGGCAGCAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACGAAGCAGAUAUGU ....(((((((((.(((((.....)))))..)))................((.(.(((((...((((((((((.(((.....))).))))))))))))))).).))...))))))..... ( -32.10, z-score = -2.04, R) >droEre2.scaffold_4784 2724675 91 - 25762168 -----------------------------GCUACAACAAGAUUUGGUUGAUUUCCUUGCAAAUUGGCAGCACAUGUUUGCUCGACCAGUGCUGCCAAGCUCAGCUACAAAGCAGAAUUGC -----------------------------....((((........))))........((((.((((((((((.((.((....)).)))))))))))).....(((....)))....)))) ( -25.70, z-score = -0.62, R) >droYak2.chr3L 14965046 111 + 24197627 ---------GCCCACUCUUGUGUUUAUGUCCUUUAGCAAGAUUUGAUUGAUUUCCUUGCAAAUUGGCAUCACUUGUUUGCCCGACCAGUGCUGCCAAGCUAGGCUACAAAGCAGAUCUGU ---------...(((....))).............(((.((((((........(((.((...((((((.((((.(((.....))).)))).)))))))).)))........))))))))) ( -27.89, z-score = -0.76, R) >droSec1.super_2 2726035 120 - 7591821 GUUUUCUGCUGUACAGAAAAGUUCUUUCUUCUACAGUAACAUUUGAAAAAUGUCCUUGCUAAUUGGCAGGAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACAAAGCAGAUCUGU ......(((((((.(((((.....)))))..)))))))..(((((.....((.(.(((((...((((((.(((.(((.....))).))).))))))))))).).)).....))))).... ( -31.20, z-score = -1.20, R) >droSim1.chr3L 2256364 104 - 22553184 ----------------AAAAUUUCUUUAUUCUACAGUAACAUUUGAAUAAAGUCCUUGCUAAUUGGCAGCAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACAAAGCAGAUCUGA ----------------.......((((((((.............))))))))...(((((...((((((((((.(((.....))).)))))))))))))))................... ( -27.42, z-score = -2.20, R) >consensus ________________AAAAUUUCUUUCUUCUACAGUAACAUUUGAAUAAUGUCCUUGCUAAUUGGCAGCAUUUGUUUGCUCGACCAGUGCUGCCAAGCAAUGUCACAAAGCAGAUCUGU .................................(((....(((((........(.((((...(((((((((((.(((.....))).))))))))))))))).)........)))))))). (-19.23 = -20.19 + 0.96)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:58:14 2011