
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 2,432,492 – 2,432,609 |
| Length | 117 |
| Max. P | 0.994716 |

| Location | 2,432,492 – 2,432,609 |
|---|---|
| Length | 117 |
| Sequences | 7 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 63.42 |
| Shannon entropy | 0.66504 |
| G+C content | 0.43150 |
| Mean single sequence MFE | -29.94 |
| Consensus MFE | -15.14 |
| Energy contribution | -13.89 |
| Covariance contribution | -1.26 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.73 |
| SVM RNA-class probability | 0.994716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 2432492 117 + 24543557 UCCCGGAAAGGAAAUUGGCUAGCGA--UAAGCACGAGAGGUGCGCAAGUCUUGUGUUUAUCUUUGCGUUCAAUAUUCAUUCUUUUAUAUAUGUACAUUCUCCGUUUUACAGCCGGGUAA- .(((((((((((((((((...((((--((((((((((....((....))))))))))))))...))..))))).....))))))).....((((.(........).)))).)))))...- ( -32.90, z-score = -2.16, R) >droEre2.scaffold_4784 2448416 109 + 25762168 UCCCGGAAAGGAAAUUUGCUAGCGA--UAAGCGCGAGAGGUGUGCAUGUCUUGUGUUUAUCUUUGAGUUCAAUAUUCUUU--------UAUACACAUUCUCCGGUUUACAGCCGGCGAA- .....(((((((.(((.(((...((--(((((((((((.((....)).)))))))))))))....)))..))).))))))--------).......(((.(((((.....))))).)))- ( -34.90, z-score = -2.82, R) >droYak2.chr3L 14695164 110 - 24197627 UCCCGGAAAGGAAAUUUGCUAGCGA--UAAGCACGAGAGGUGUGCAAGUCUUGUGUUUAUCAUUGAGUUCAAUAUUCUCU--------UAUACACAUUCGCCGUUUUACGACCGGGAAAA ((((((....(((((..((....((--(((((((((((..(....)..)))))))))))))((((....)))).......--------...........)).)))))....))))))... ( -33.10, z-score = -2.69, R) >droSec1.super_2 2463627 114 + 7591821 UCCCGGAAAGGAAAUUGGCUAGCGA--UAAGCACGAGCGGUGUGCAAGUCUUGUGUUUAUCUUUGCGUUCAAUAUUCAUUCUUU----UAUAUACAUUCCUUGUUUUACAGUCGGGGGAA ((((((((.(((.(((((...((((--(((((((((((.........).))))))))))))...))..))))).))).))))..----..........(((((.....)))..)))))). ( -29.30, z-score = -1.07, R) >droSim1.chr3L 1995168 116 + 22553184 UCCCGGAAAGGAAAUUGGCUAGCGAGAUAAGCACGAGAGGUGUGCAAUUCUUGUGUUUAUCUUUGCGUUCAAUAUUCAUUCUUU----UAUAUACAUUCCUUGUUUCACAGUCGGGGGAA ((((((((.(((.(((((...((((((((((((((((((........)))))))))))))).))))..))))).))).))))..----..........(((((.....)))..)))))). ( -36.40, z-score = -3.18, R) >dp4.chrXR_group6 12398838 106 + 13314419 -UCCGGCUGAAAAGUGUUUUUGUUU--UGA---CGAAAAUUCAUUUUGUUUUGAGUUCAACGAAACGGGAAGUUUCCUCCCCU-----CC--CCUCCUCCCCCUCCAACAAUUCAUGCA- -....((((((...((((...((((--((.---.((..(((((........)))))))..))))))((((.......))))..-----..--..............)))).)))).)).- ( -19.40, z-score = -0.25, R) >droPer1.super_33 419829 108 + 967471 -UCCGGCUGAAAAGUGUUUUUGUUU--UGA---CGAAAAUUCAUUUUGUUUUGAGUUCAACGAAACGGUAAGUGUCCCCCCCU-----CCGUCCCUCCCCCCCUCAAACAGCUCGGGCC- -((((((((.(((((((((((((..--..)---))))))..))))))..((((((.........((((...............-----))))..........)))))))))).))))..- ( -23.57, z-score = -0.75, R) >consensus UCCCGGAAAGGAAAUUUGCUAGCGA__UAAGCACGAGAGGUGUGCAAGUCUUGUGUUUAUCUUUGCGUUCAAUAUUCAUUCUU_____UAUACACAUUCCCCGUUUUACAGCCGGGGAA_ .(((((..((........)).((((..(((((((((((..........)))))))))))...)))).............................................))))).... (-15.14 = -13.89 + -1.26)

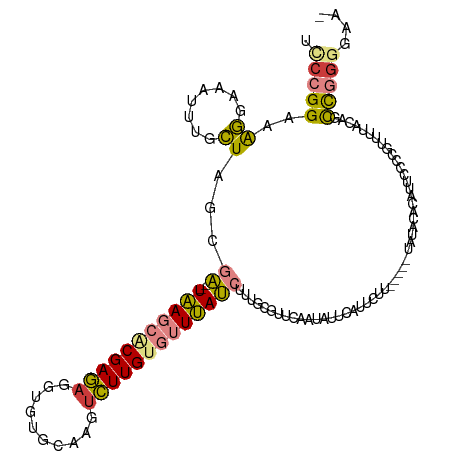
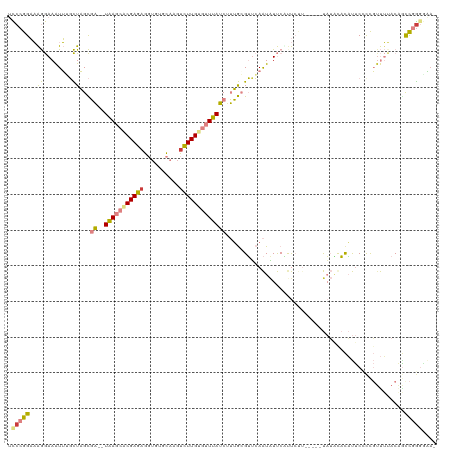
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:57:39 2011