
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 2,380,587 – 2,380,683 |
| Length | 96 |
| Max. P | 0.917322 |

| Location | 2,380,587 – 2,380,683 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 61.97 |
| Shannon entropy | 0.72947 |
| G+C content | 0.43613 |
| Mean single sequence MFE | -28.04 |
| Consensus MFE | -9.42 |
| Energy contribution | -10.26 |
| Covariance contribution | 0.85 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.917322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
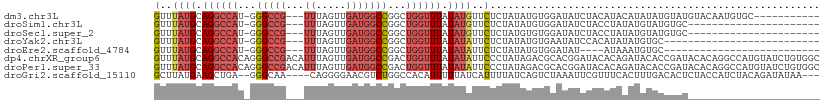
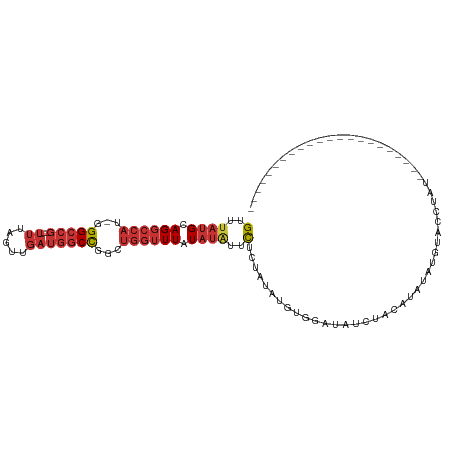
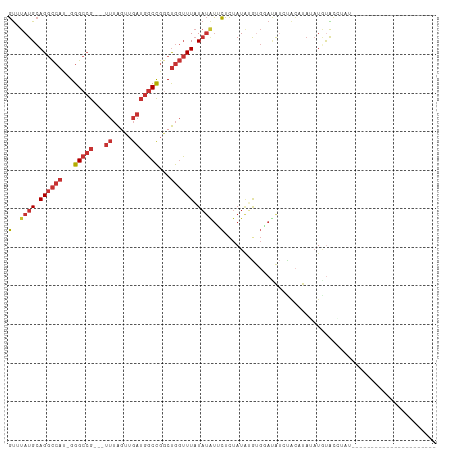
>dm3.chr3L 2380587 96 + 24543557 GUUUAUGCAGGCCAU-GGGCCG---UUUAGUUGAUGGCCGGCUGGUUUAUAUGUUCUCUAUAUGUGGAUAUCUACAUACAUAUAUGUAUGUACAAUGUGC----------- ((.((((((((((((-.(((((---((.....))))))).).)))))(((((((.......((((((....))))))))))))))))))).)).......----------- ( -30.91, z-score = -1.69, R) >droSim1.chr3L 1942690 85 + 22553184 GUUUAUGCAGGCCAU-GGGCCG---UUUAGUUGAUGGCCGGCUGGUUUAUAUGUUCUCUAUAUGUGGAUAUCUACCUAUAUGUAUGUGC---------------------- (..((((.(((((((-.(((((---((.....))))))).).)))))).))))..)..(((((((((........))))))))).....---------------------- ( -25.90, z-score = -1.21, R) >droSec1.super_2 2405169 85 + 7591821 GUUUAUGCAGGCCAU-GGGCCG---UUUAGUUGAUGGCCGGCUGGUUUAUAUGUUCUCUAUGUGUGGAUAUCUACCUAUAUGUAUGUGC---------------------- ((.((((((((((..-.(((((---((.....)))))))))))(((....((((((.(.....).))))))..)))....)))))).))---------------------- ( -25.90, z-score = -1.16, R) >droYak2.chr3L 14641792 81 - 24197627 GUUUAUGCAGGCCAU-GGGCCG---UUUAGUUGAUGGCCGGCUGGUUUAUAUAUUCUCUAUAUGUGAAUAUCCACAUAUAUGUGC-------------------------- (..((((.(((((((-.(((((---((.....))))))).).)))))).))))..)..((((((((......)))))))).....-------------------------- ( -27.30, z-score = -2.28, R) >droEre2.scaffold_4784 2395822 77 + 25762168 GUUUAUGCAGGCCAU-GGGCCG---UUUAGUUGAUGGCCGGCUGGUUUAUAUAUUCUCUAUAUGUGGAUAU----AUAAAUGUGC-------------------------- ......(((((((..-.(((((---((.....))))))))))).((((((((((((.(.....).))))))----)))))).)))-------------------------- ( -27.30, z-score = -2.95, R) >dp4.chrXR_group6 12332008 111 + 13314419 GUUUAUGCAGGCCACAGGGCCGACAUUUAGUUGAUGGCCGACUGGUUUAUAUAUUCCCUAUAGACGCACGGAUACACAGAUACACCGAUACACAGGCCAUGUAUCUGUGGC (((..(((.((((....))))......((((((.....))))))(((((((.......))))))))))..))).((((((((((((........))...)))))))))).. ( -37.30, z-score = -3.07, R) >droPer1.super_33 353255 111 + 967471 GUUUAUGCAGGCCACAGGGCCGACAUUUAGUUGAUGGCCGACUGGUUUAUAUAUUCCCUAUAGACGCACGGAUACACAGAUACACCGAUACACAGGCCAUGUAUCUGUGGC (((..(((.((((....))))......((((((.....))))))(((((((.......))))))))))..))).((((((((((((........))...)))))))))).. ( -37.30, z-score = -3.07, R) >droGri2.scaffold_15110 24215054 102 - 24565398 GCUUAUGAAGCUGA--GGGCAA----CAGGGGAACGUCUGGCCACAUUUUUAUCAUUUUAUCAGUCUAAAUUCGUUUCACUUUGACACUCUACCAUCUACAGAUAUAA--- (((.....))).((--(..(((----.((.((((((...(((.....................)))......)))))).)))))...)))....(((....)))....--- ( -12.40, z-score = 1.64, R) >consensus GUUUAUGCAGGCCAU_GGGCCG___UUUAGUUGAUGGCCGGCUGGUUUAUAUAUUCUCUAUAUGUGGAUAUCUACAUAUAUGUACCUAU______________________ (..((((.((((((...(((((............)))))...)))))).))))..)....................................................... ( -9.42 = -10.26 + 0.85)


| Location | 2,380,587 – 2,380,683 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 61.97 |
| Shannon entropy | 0.72947 |
| G+C content | 0.43613 |
| Mean single sequence MFE | -20.78 |
| Consensus MFE | -8.27 |
| Energy contribution | -8.73 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.800277 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
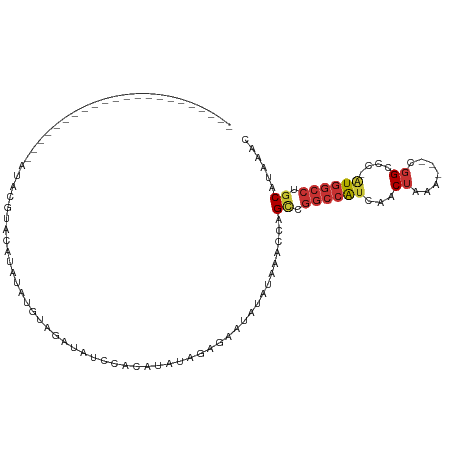
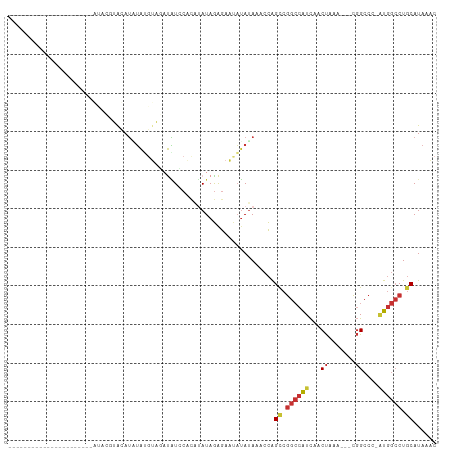
>dm3.chr3L 2380587 96 - 24543557 -----------GCACAUUGUACAUACAUAUAUGUAUGUAGAUAUCCACAUAUAGAGAACAUAUAAACCAGCCGGCCAUCAACUAAA---CGGCCC-AUGGCCUGCAUAAAC -----------......(.((((((((....)))))))).)............................((.((((((...((...---.))...-)))))).))...... ( -19.90, z-score = -1.37, R) >droSim1.chr3L 1942690 85 - 22553184 ----------------------GCACAUACAUAUAGGUAGAUAUCCACAUAUAGAGAACAUAUAAACCAGCCGGCCAUCAACUAAA---CGGCCC-AUGGCCUGCAUAAAC ----------------------.............(((..((((...............))))..))).((.((((((...((...---.))...-)))))).))...... ( -14.56, z-score = -0.82, R) >droSec1.super_2 2405169 85 - 7591821 ----------------------GCACAUACAUAUAGGUAGAUAUCCACACAUAGAGAACAUAUAAACCAGCCGGCCAUCAACUAAA---CGGCCC-AUGGCCUGCAUAAAC ----------------------.............(((..((((...............))))..))).((.((((((...((...---.))...-)))))).))...... ( -14.56, z-score = -0.95, R) >droYak2.chr3L 14641792 81 + 24197627 --------------------------GCACAUAUAUGUGGAUAUUCACAUAUAGAGAAUAUAUAAACCAGCCGGCCAUCAACUAAA---CGGCCC-AUGGCCUGCAUAAAC --------------------------.....(((((((((....)))))))))................((.((((((...((...---.))...-)))))).))...... ( -20.20, z-score = -2.41, R) >droEre2.scaffold_4784 2395822 77 - 25762168 --------------------------GCACAUUUAU----AUAUCCACAUAUAGAGAAUAUAUAAACCAGCCGGCCAUCAACUAAA---CGGCCC-AUGGCCUGCAUAAAC --------------------------.....(((((----((((.(.(.....).).)))))))))...((.((((((...((...---.))...-)))))).))...... ( -15.90, z-score = -2.47, R) >dp4.chrXR_group6 12332008 111 - 13314419 GCCACAGAUACAUGGCCUGUGUAUCGGUGUAUCUGUGUAUCCGUGCGUCUAUAGGGAAUAUAUAAACCAGUCGGCCAUCAACUAAAUGUCGGCCCUGUGGCCUGCAUAAAC (((((((((((((.....)))))))(((.....(((((((((.((......)).)).))))))).)))....((((..((......))..)))).)))))).......... ( -33.90, z-score = -1.12, R) >droPer1.super_33 353255 111 - 967471 GCCACAGAUACAUGGCCUGUGUAUCGGUGUAUCUGUGUAUCCGUGCGUCUAUAGGGAAUAUAUAAACCAGUCGGCCAUCAACUAAAUGUCGGCCCUGUGGCCUGCAUAAAC (((((((((((((.....)))))))(((.....(((((((((.((......)).)).))))))).)))....((((..((......))..)))).)))))).......... ( -33.90, z-score = -1.12, R) >droGri2.scaffold_15110 24215054 102 + 24565398 ---UUAUAUCUGUAGAUGGUAGAGUGUCAAAGUGAAACGAAUUUAGACUGAUAAAAUGAUAAAAAUGUGGCCAGACGUUCCCCUG----UUGCCC--UCAGCUUCAUAAGC ---...........((.(((.(((.(((.((((.......)))).)))....................((((((........)))----..))))--)).)))))...... ( -13.30, z-score = 1.36, R) >consensus ______________________AUACGUACAUAUAUGUAGAUAUCCACAUAUAGAGAAUAUAUAAACCAGCCGGCCAUCAACUAAA___CGGCCC_AUGGCCUGCAUAAAC .....................................................................((.((((((...((.......))....)))))).))...... ( -8.27 = -8.73 + 0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:57:35 2011