
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 2,192,030 – 2,192,122 |
| Length | 92 |
| Max. P | 0.579415 |

| Location | 2,192,030 – 2,192,122 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 76.43 |
| Shannon entropy | 0.46795 |
| G+C content | 0.33016 |
| Mean single sequence MFE | -18.75 |
| Consensus MFE | -7.04 |
| Energy contribution | -7.16 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.579415 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
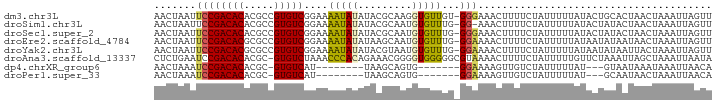
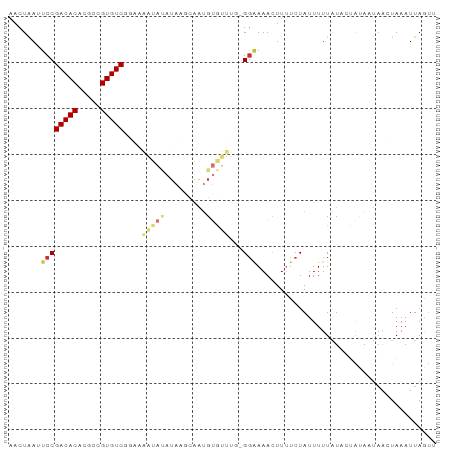
>dm3.chr3L 2192030 92 + 24543557 AACUAAUUCCGACACACGCCGUGUCGGAAAAUAUAUACGCAAGGUGUUGU-GGGAAACUUUUCUAUUUUUAUACUGCACUAACUAAAUUAGUU (((((((((((((((.....))))))))).........(((..(((..((-((((.....))))))...)))..)))..........)))))) ( -25.10, z-score = -3.59, R) >droSim1.chr3L 1748334 91 + 22553184 AACUAAUUCCGACACACGCCGUGUCGGAAAAUAUAUACGCAAUGUGUUUG-GG-AAACUUUUCUAUUUUUAUACUAUACUAACUAAAUUAGUU (((((((((((((((.....))))))))((((((((.....))))))))(-(.-...))...........................))))))) ( -21.10, z-score = -3.38, R) >droSec1.super_2 2218067 92 + 7591821 AACUAAUUCCGACACACGCCGUGUCGGAAAAUAUAUACGCAAUGUGUUUG-GGGAAACUUUUCUAUUUUUAUACUAUACUAACUAAAUUAGUU (((((((((((((((.....))))))))((((((((.....))))))))(-((....)))..........................))))))) ( -21.60, z-score = -3.17, R) >droEre2.scaffold_4784 2186716 92 + 25762168 AACUAAUUCCGACACGCGCCGUGUCGGAAAAUAUAUAAGCAAUGUGUUUG-GGAAAACUUUUCUAUUUUUAUAAUAUAAUAACUAAAUUAGUU ((((((((((((((((...)))))))))((((((((.....))))))))(-((((....)))))......................))))))) ( -21.10, z-score = -2.92, R) >droYak2.chr3L 2145361 92 + 24197627 AACUAAUUCCGACACGCGCCGUGUCGGAAAAUAUAUACGUAAUGUGUUUG-GGAAAACUUUUCUAUUUUUAUAAUAUAAUUACUAAAUUAGUU ((((((((((((((((...)))))))))).........((((((((((..-.(((((........)))))..))))).)))))....)))))) ( -23.40, z-score = -3.44, R) >droAna3.scaffold_13337 22649100 92 - 23293914 CUCUGAAUCCGACACACGC-GUGUCUAAACCCACAGAAACGGGGUGGGGGCGUAAAACUUUUCUAUUUUUGUUCUAAAUUAGCUAAAUUAAUA ..((((..(((((((....-)))))...((((.(......)))))))(((((.((((........)))))))))....))))........... ( -15.70, z-score = 0.51, R) >dp4.chrXR_group6 12103238 74 + 13314419 AACUAAAUCCGACACACGC-GUGUCAU--------UAAGCAGUG-------GGAAAAGUUGUCUAUUUUUAU---GUAAUAAAUAAAUUAACA ..........(((((....-)))))((--------((....(((-------((((.((....)).)))))))---.))))............. ( -10.00, z-score = -0.02, R) >droPer1.super_33 126212 74 + 967471 AACUAAAUCCGACACACGC-GUGUCAU--------UAAGCAGUG-------GGAAAAGUUGUCUAUUUUUAU---GCAAUAACUAAAUUAACA ..........(((((....-)))))..--------...(((.((-------((((.((....)).)))))))---))................ ( -12.00, z-score = -0.65, R) >consensus AACUAAUUCCGACACACGCCGUGUCGGAAAAUAUAUAAGCAAUGUGUUUG_GGAAAACUUUUCUAUUUUUAUACUAUAAUAACUAAAUUAGUU .......((((((((.....)))))....(((((.........)))))...)))....................................... ( -7.04 = -7.16 + 0.13)

| Location | 2,192,030 – 2,192,122 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 76.43 |
| Shannon entropy | 0.46795 |
| G+C content | 0.33016 |
| Mean single sequence MFE | -15.51 |
| Consensus MFE | -9.62 |
| Energy contribution | -10.01 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.25 |
| Mean z-score | -0.98 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.521816 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
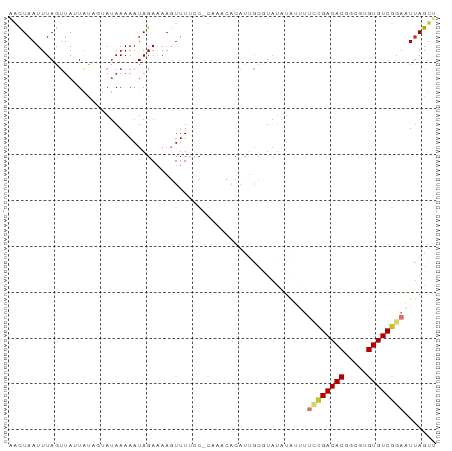
>dm3.chr3L 2192030 92 - 24543557 AACUAAUUUAGUUAGUGCAGUAUAAAAAUAGAAAAGUUUCCC-ACAACACCUUGCGUAUAUAUUUUCCGACACGGCGUGUGUCGGAAUUAGUU ((((((..........((((..........(((....)))..-........)))).........(((((((((.....))))))))))))))) ( -18.25, z-score = -1.16, R) >droSim1.chr3L 1748334 91 - 22553184 AACUAAUUUAGUUAGUAUAGUAUAAAAAUAGAAAAGUUU-CC-CAAACACAUUGCGUAUAUAUUUUCCGACACGGCGUGUGUCGGAAUUAGUU ((((((........((((((((........(((....))-).-.........))).)))))...(((((((((.....))))))))))))))) ( -17.67, z-score = -1.59, R) >droSec1.super_2 2218067 92 - 7591821 AACUAAUUUAGUUAGUAUAGUAUAAAAAUAGAAAAGUUUCCC-CAAACACAUUGCGUAUAUAUUUUCCGACACGGCGUGUGUCGGAAUUAGUU ((((((........((((((((....................-.........))).)))))...(((((((((.....))))))))))))))) ( -17.25, z-score = -1.43, R) >droEre2.scaffold_4784 2186716 92 - 25762168 AACUAAUUUAGUUAUUAUAUUAUAAAAAUAGAAAAGUUUUCC-CAAACACAUUGCUUAUAUAUUUUCCGACACGGCGCGUGUCGGAAUUAGUU ((((((((................((((((...((((.....-..........))))...))))))((((((((...)))))))))))))))) ( -18.86, z-score = -2.52, R) >droYak2.chr3L 2145361 92 - 24197627 AACUAAUUUAGUAAUUAUAUUAUAAAAAUAGAAAAGUUUUCC-CAAACACAUUACGUAUAUAUUUUCCGACACGGCGCGUGUCGGAAUUAGUU ((((((....(((((.........(((((......)))))..-.......))))).........((((((((((...)))))))))))))))) ( -19.67, z-score = -2.74, R) >droAna3.scaffold_13337 22649100 92 + 23293914 UAUUAAUUUAGCUAAUUUAGAACAAAAAUAGAAAAGUUUUACGCCCCCACCCCGUUUCUGUGGGUUUAGACAC-GCGUGUGUCGGAUUCAGAG ....(((((..(((.(((......))).)))..))))).....(((((((.........)))))....(((((-....)))))))........ ( -15.10, z-score = 0.25, R) >dp4.chrXR_group6 12103238 74 - 13314419 UGUUAAUUUAUUUAUUAC---AUAAAAAUAGACAACUUUUCC-------CACUGCUUA--------AUGACAC-GCGUGUGUCGGAUUUAGUU ((((.((((.........---....)))).))))........-------.((((....--------.((((((-....))))))....)))). ( -8.62, z-score = 0.34, R) >droPer1.super_33 126212 74 - 967471 UGUUAAUUUAGUUAUUGC---AUAAAAAUAGACAACUUUUCC-------CACUGCUUA--------AUGACAC-GCGUGUGUCGGAUUUAGUU ((((.((((.........---....)))).))))........-------.((((....--------.((((((-....))))))....)))). ( -8.62, z-score = 1.04, R) >consensus AACUAAUUUAGUUAUUAUAGUAUAAAAAUAGAAAAGUUUUCC_CAAACACAUUGCGUAUAUAUUUUCCGACACGGCGUGUGUCGGAAUUAGUU .................................................................((((((((.....))))))))....... ( -9.62 = -10.01 + 0.39)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:57:12 2011