
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 2,149,714 – 2,149,813 |
| Length | 99 |
| Max. P | 0.924950 |

| Location | 2,149,714 – 2,149,813 |
|---|---|
| Length | 99 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 69.66 |
| Shannon entropy | 0.69588 |
| G+C content | 0.54509 |
| Mean single sequence MFE | -31.88 |
| Consensus MFE | -17.85 |
| Energy contribution | -18.43 |
| Covariance contribution | 0.57 |
| Combinations/Pair | 1.37 |
| Mean z-score | -0.76 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.924950 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
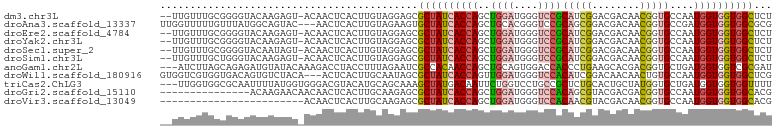
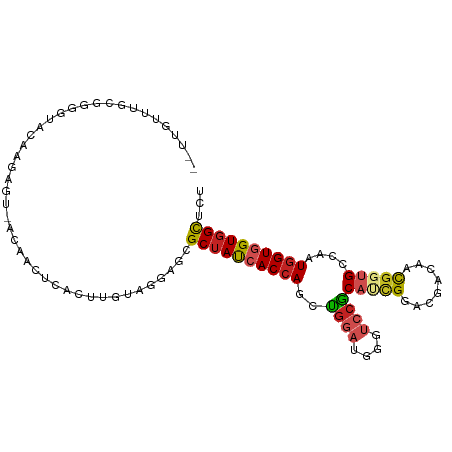
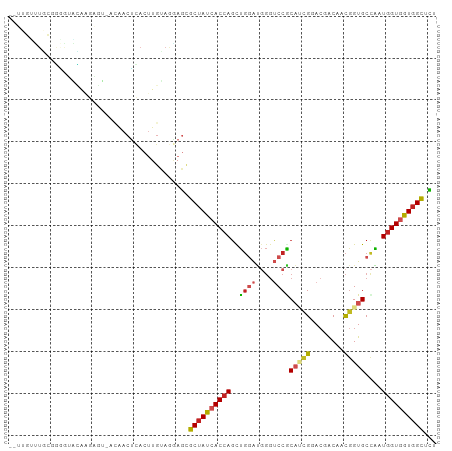
>dm3.chr3L 2149714 99 + 24543557 --UUGUUUGCGGGGUACAAGAGU-ACAACUCACUUGUAGGAGCGCUAUCACCAGCUGGAUGGGUCCGCAUCGGACGACAACGGUGCCAAUGGUGGUGGCUCU --......((....((((((((.-....)))..)))))...))((((((((((...(((....)))((((((........))))))...))))))))))... ( -34.50, z-score = -0.89, R) >droAna3.scaffold_13337 22611035 99 - 23293914 UUGGUUUUUGUUUAUGGCAGUAC---AACUCACUUGUAGAAGUGCUAUCACCAGCUGCACGGGUCCGCAGCGGACGACAACGGUGCCGAUGGUGGUGGCGCG .......((((.....))))(((---((.....)))))...((((((((((((.(.((((.((((((...))))).....).)))).).)))))))))))). ( -35.20, z-score = -0.82, R) >droEre2.scaffold_4784 2144582 99 + 25762168 --UUGUUUGCGGGGUACAAGAGU-ACAACUCACUUGUAGGAGCGCUAUCACCAGCUGGAUGGGUCCGCAUCGGACGACAACGGUGCCAAUGGUGGUGGCUCU --......((....((((((((.-....)))..)))))...))((((((((((...(((....)))((((((........))))))...))))))))))... ( -34.50, z-score = -0.89, R) >droYak2.chr3L 2102712 99 + 24197627 --UUGUUUGCGGGGUACAAGAGU-ACAACUCACUUGUAGGAGCGCUAUCACCAGCUGGAUGGGUCCGCAUCGGACGACAACGGUGCCAAUGGUGGUGGCUCU --......((....((((((((.-....)))..)))))...))((((((((((...(((....)))((((((........))))))...))))))))))... ( -34.50, z-score = -0.89, R) >droSec1.super_2 2175841 99 + 7591821 --UUGUUUGCGGGGUACAAUAGU-ACAACUCACUUGUAGGAGCGCUAUCACCAGCUGGAUGGGUCCGCAUCGGACGACAACGGUGCCAAUGGUGGUGGCUCU --...((((((((((((....))-))......))))))))((.((((((((((...(((....)))((((((........))))))...)))))))))).)) ( -34.50, z-score = -1.08, R) >droSim1.chr3L 1705539 99 + 22553184 --UUGUUUGCUGGGUACAAGAGU-ACAACUCACUUGUAGGAGCGCUAUCACCAGCUGGAUGGGUCCGCAUCGGACGACAACGGUGCCAAUGGUGGUGGCUCU --......(((...((((((((.-....)))..)))))..)))((((((((((...(((....)))((((((........))))))...))))))))))... ( -35.20, z-score = -1.14, R) >anoGam1.chr2L 1786196 99 - 48795086 ---AUCUUAGCAGAGAUGUAUACAAAGACCUACCUUUAGAAUCGCCACAACCAGCUGCAGUGGACCACCCUGAAGCACGACGGUGCUGAUGGUGGUCGCGAU ---(((((....)))))......((((......))))...((((((((...........)))(((((((....(((((....)))))...)))))))))))) ( -29.60, z-score = -1.72, R) >droWil1.scaffold_180916 1457577 99 - 2700594 GUGGUCGUGGUGACAGUGUCUACA---ACUCACUUGCAAUAGCGCUAUCACCAGUUGGAUGGGUCCACAUCGGACAACAACUGUGCCAAUGGUGGUGGCUCG ((((.((((....)).)).)))).---........((....))((((((((((.((((((((((((.....)))).....)))).))))))))))))))... ( -35.40, z-score = -1.17, R) >triCas2.ChLG3 2759232 99 - 32080666 ---UUGGUGGCGCAAUUUUAUGGUGGGACGUACAUGCAGCAAAGCUAUGACAAUUCUGGUCCUGCCGCUCUGCACUGCUAUGGUGCUGAUGGUGGUGGUUUU ---...(..(.((........((..((((...(((((......)).))).((....))))))..)))).)..)((..((((.((....)).))))..))... ( -26.80, z-score = 1.20, R) >droGri2.scaffold_15110 13161083 87 - 24565398 ---------------ACAAGAACAACAACUCACUUGCAAGAGCGCUAUCACCAGCUGGAUGGGUCCACAGCGUACGACGACGGUGCCAAUGGUGGUGGCACG ---------------.............(((........))).((((((((((((((..((....))))))((((.......))))...))))))))))... ( -26.50, z-score = -0.51, R) >droVir3.scaffold_13049 13267935 78 - 25233164 ------------------------ACAACUCACUUGCAAGAGCGCUAUCACCAGCUGGAUGGGUCCACAACGUACGACAACGGUGCCAAUGGUGGUGGCACG ------------------------....(((........))).(((((((((((.((((....)))))...((((.......))))...))))))))))... ( -24.00, z-score = -0.45, R) >consensus __UUGUUUGCGGGGUACAAGAGU_ACAACUCACUUGUAGGAGCGCUAUCACCAGCUGGAUGGGUCCGCAUCGGACGACAACGGUGCCAAUGGUGGUGGCUCU ...........................................((((((((((..((((....))))(((((........)))))....))))))))))... (-17.85 = -18.43 + 0.57)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:57:09 2011