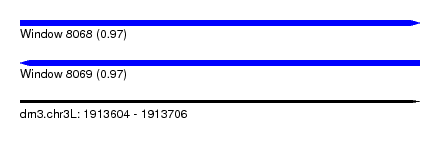

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 1,913,604 – 1,913,706 |
| Length | 102 |
| Max. P | 0.971249 |

| Location | 1,913,604 – 1,913,706 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 79.04 |
| Shannon entropy | 0.36514 |
| G+C content | 0.36426 |
| Mean single sequence MFE | -23.54 |
| Consensus MFE | -13.74 |
| Energy contribution | -15.18 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.969437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
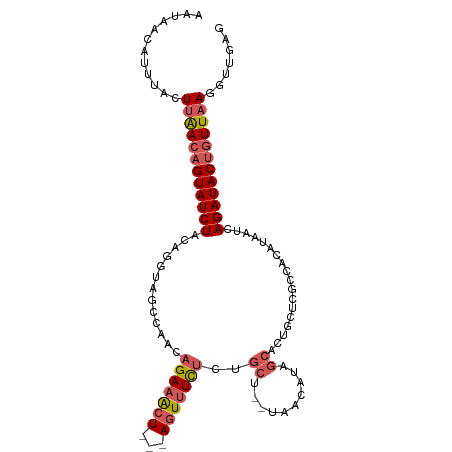
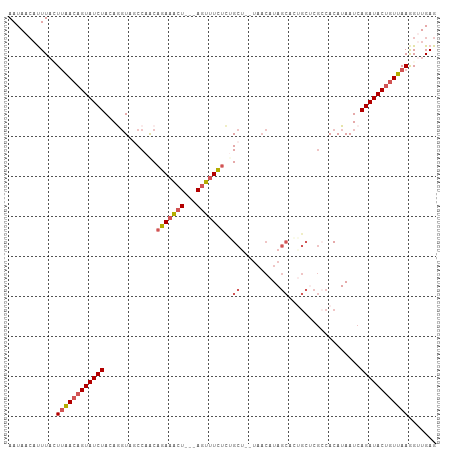
>dm3.chr3L 1913604 102 + 24543557 AAUAACAUUUACUUAACAGUAUCUACAGGUAGCCAACAGAAACU---AGUUUCUCUGCU--UAACAUAGCAAGGCCCGACACAUAAUCAGAUACUGUUAAGGUUGAG ...........(((((((((((((...((..(((...(((((..---..))))).((((--......)))).)))))((.......)))))))))))))))...... ( -28.90, z-score = -4.37, R) >droEre2.scaffold_4784 1902339 107 + 25762168 AAUAACAUUUAUUUGACAGUAUCUACAAGUUUCCAGCAGAAACUGCAAUUAUUGUUGCUGCUAACAUAGCACUGCUCGCCACAUAAUCAGAUACUGUUAAAGCUGAG ...........(((((((((((((....((..(.(((((.....(((((....)))))(((((...)))))))))).)..))......)))))))))))))...... ( -28.90, z-score = -3.24, R) >droYak2.chr3L 1867550 85 + 24197627 AAUACAUCUUUCUUGAAUGUAUCUACAACUUUCCAACAGAAACU---AGUUUCUCUGCU-------UA------------ACAUAAUCAGAUACUGUUAAAGUUGAG .(((((((......).))))))...(((((((..((((((((..---..))))((((.(-------((------------...))).))))...))))))))))).. ( -14.00, z-score = -1.40, R) >droSec1.super_2 1939310 101 + 7591821 AAUAACAUUUAGU-AACAGUAUCUACAGGUAGCCAACAGAAGCU---AGUUUCUCGGCA--UAACAUAGCACUGCUCGCCACAUAAUCAGAUACUGUUAAGGUUGAG ..((((......(-((((((((((....((.((...(((..(((---((((........--.))).)))).)))...)).))......)))))))))))..)))).. ( -23.50, z-score = -1.89, R) >droSim1.chr3L 1468981 102 + 22553184 AAUAACAUUUAGUUAACAGUAUCUACAGGUAGCCAACAGAAACU---AGUUUCUCGGCA--UAACAUAGCACUGCUCGCCACAUAAUCAGAUACUGUUAAGGUUGAG ............((((((((((((.......(((...(((((..---..))))).))).--.......((.......)).........))))))))))))....... ( -22.40, z-score = -1.96, R) >consensus AAUAACAUUUACUUAACAGUAUCUACAGGUAGCCAACAGAAACU___AGUUUCUCUGCU__UAACAUAGCACUGCUCGCCACAUAAUCAGAUACUGUUAAGGUUGAG ............((((((((((((.............((((((.....))))))..(((........)))..................))))))))))))....... (-13.74 = -15.18 + 1.44)

| Location | 1,913,604 – 1,913,706 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 79.04 |
| Shannon entropy | 0.36514 |
| G+C content | 0.36426 |
| Mean single sequence MFE | -29.10 |
| Consensus MFE | -14.47 |
| Energy contribution | -17.54 |
| Covariance contribution | 3.07 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.971249 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
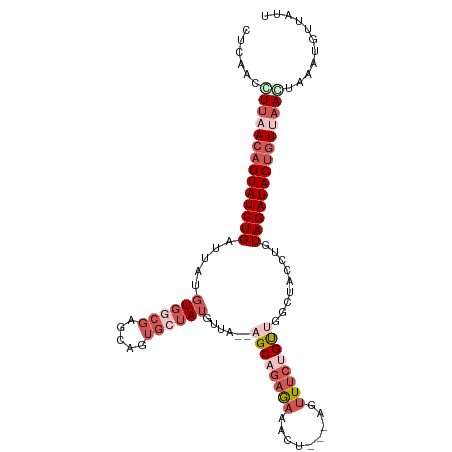
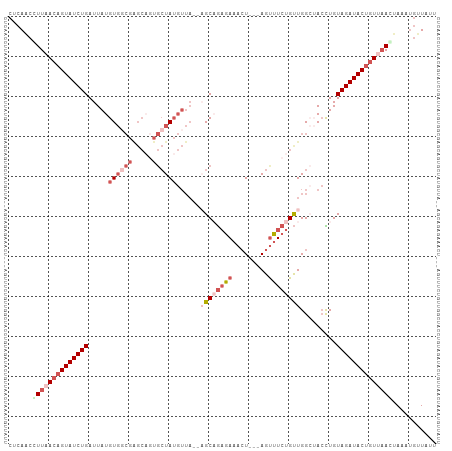
>dm3.chr3L 1913604 102 - 24543557 CUCAACCUUAACAGUAUCUGAUUAUGUGUCGGGCCUUGCUAUGUUA--AGCAGAGAAACU---AGUUUCUGUUGGCUACCUGUAGAUACUGUUAAGUAAAUGUUAUU ......((((((((((((((((.....))).(((((((((......--)))))(((((..---..)))))...))))......)))))))))))))........... ( -30.60, z-score = -3.22, R) >droEre2.scaffold_4784 1902339 107 - 25762168 CUCAGCUUUAACAGUAUCUGAUUAUGUGGCGAGCAGUGCUAUGUUAGCAGCAACAAUAAUUGCAGUUUCUGCUGGAAACUUGUAGAUACUGUCAAAUAAAUGUUAUU ...(((((((((((((((((.....((..(.((((((((((...)))))((((......)))).....))))).)..))...))))))))))....)))).)))... ( -30.20, z-score = -1.77, R) >droYak2.chr3L 1867550 85 - 24197627 CUCAACUUUAACAGUAUCUGAUUAUGU------------UA-------AGCAGAGAAACU---AGUUUCUGUUGGAAAGUUGUAGAUACAUUCAAGAAAGAUGUAUU (((((((((..(((...))).......------------..-------((((((((....---..))))))))..))))))).))(((((((.......))))))). ( -17.40, z-score = -0.98, R) >droSec1.super_2 1939310 101 - 7591821 CUCAACCUUAACAGUAUCUGAUUAUGUGGCGAGCAGUGCUAUGUUA--UGCCGAGAAACU---AGCUUCUGUUGGCUACCUGUAGAUACUGUU-ACUAAAUGUUAUU ........((((((((((((.....(((((.(((((.((((.(((.--........))))---)))..))))).)))))...)))))))))))-)............ ( -34.10, z-score = -4.32, R) >droSim1.chr3L 1468981 102 - 22553184 CUCAACCUUAACAGUAUCUGAUUAUGUGGCGAGCAGUGCUAUGUUA--UGCCGAGAAACU---AGUUUCUGUUGGCUACCUGUAGAUACUGUUAACUAAAUGUUAUU .......(((((((((((((.....(((((.(((((.((((.(((.--........))))---)))..))))).)))))...)))))))))))))............ ( -33.20, z-score = -4.21, R) >consensus CUCAACCUUAACAGUAUCUGAUUAUGUGGCGAGCAGUGCUAUGUUA__AGCAGAGAAACU___AGUUUCUGUUGGCUACCUGUAGAUACUGUUAACUAAAUGUUAUU ......((((((((((((((.....((((((.....))))))......((((((((.........)))))))).........))))))))))))))........... (-14.47 = -17.54 + 3.07)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:56:43 2011