| Sequence ID | dm3.chr3L |
|---|---|
| Location | 1,683,780 – 1,683,874 |
| Length | 94 |
| Max. P | 0.984987 |

| Location | 1,683,780 – 1,683,874 |
|---|---|
| Length | 94 |
| Sequences | 8 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 60.73 |
| Shannon entropy | 0.74293 |
| G+C content | 0.56861 |
| Mean single sequence MFE | -22.06 |
| Consensus MFE | -4.07 |
| Energy contribution | -4.58 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.25 |
| Structure conservation index | 0.18 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.984987 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 1683780 94 + 24543557 ---------AGAGACCGAACGC-GAAAUGGCAGACAAUUUGCCUAAUUCGCGUUCUGAACCCUCCAGAGCCACACCCCCUCCCCCCCUCAAGCUCCAUCCCCCC--- ---------.(((...((((((-(((..((((((...))))))...)))))))))...........(((.........))).....)))...............--- ( -23.70, z-score = -4.76, R) >droSim1.chr3L 1244608 87 + 22553184 -----AGAGAGAGACCGAACGC-GAAAUGGCAGACAAUUUGCCUAAUUCGCGUUCUGAACCCUCCAGAACC----------CCCCGCUCAA-CUCCAUCCCCCC--- -----.(((.(((.......((-(((..((((((...))))))...)))))((((((.......)))))).----------.....)))..-))).........--- ( -23.90, z-score = -4.51, R) >droSec1.super_2 1710081 87 + 7591821 ----CGAAGAGAGACCGAACGC-GAAAUGGCAGACAAUUUGCCUAAUUCGCGUUCUGAACCCUACAGAGCC----------CCC-GCUCAA-CUCCAUCCCCCC--- ----.((.(.(((...((((((-(((..((((((...))))))...)))))))))...........((((.----------...-))))..-)))).)).....--- ( -29.20, z-score = -6.11, R) >droYak2.chr3L 1635022 84 + 24197627 ---------------AGUACGC-GAAAUGGCAGACAAUUUGCCUAAUUCGCGUUCUGAACCCUACAGAGCC----CCCUCCCCCCCCUCGAAUUUCACCCCUCG--- ---------------.....((-(((..((((((...))))))...)))))((((((.......)))))).----.............................--- ( -19.30, z-score = -4.10, R) >droEre2.scaffold_4784 1667139 81 + 25762168 ---------------AAUACGC-GAAAUGGCAGACAAUUUGCCUAAUUCGCGUUCUGAGCCCUCCAGAGCC----------CCCACCUCCAGCUCCACCAGCCCCUC ---------------.....((-(((..((((((...))))))...)))))((((((.......)))))).----------..........(((.....)))..... ( -20.70, z-score = -3.36, R) >droAna3.scaffold_13337 22139161 80 - 23293914 ---------------AAAUCGC-GAAAUUGCACACAAUUUGCCUAAUUCGCGUGGAGCGGCCCCAAGCCCC--------CAUCCUCCGCCGUCCUCCUCACCGU--- ---------------.....((-(((...(((.......)))....)))))((((((.(((.....)))..--------....))))))...............--- ( -17.70, z-score = -1.41, R) >droPer1.super_27 579083 96 - 1201981 UGCAACGCGAAAGGGAGCCCACAGAAAUUGCAGACAAUUUGCCUAAUUCGCGUUCAUCGCUCUCUCAUACU------CCCUCCC-UCCCUGGCUCACCUUCUG---- ........(((.(((((((....(((...(((((...)))))....)))(((.....)))...........------.......-.....))))).)))))..---- ( -19.50, z-score = -0.78, R) >droMoj3.scaffold_6654 1003954 88 - 2564135 UGCGAGUUCAACUCG--CACACAGAAAUUGCACACAAUUUGCCUAAUUCGCGCGCGCC--UCUCUCAGGCC------CGCUCGCAUCGCCAGCUCCAA--------- ((((((.....))))--)).....((((((....))))))((...((.((.(((.(((--(.....)))).------))).)).)).)).........--------- ( -22.50, z-score = -0.98, R) >consensus _________A_AG__AGAACGC_GAAAUGGCAGACAAUUUGCCUAAUUCGCGUUCUGAACCCUCCAGAGCC________C_CCCCCCUCAAGCUCCAUCCCCCC___ ...................(((.(((...(((((...)))))....))))))....................................................... ( -4.07 = -4.58 + 0.50)
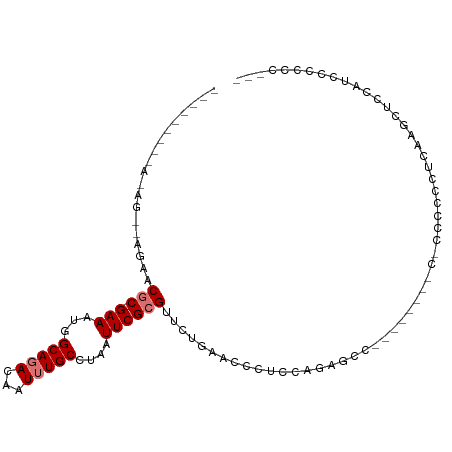


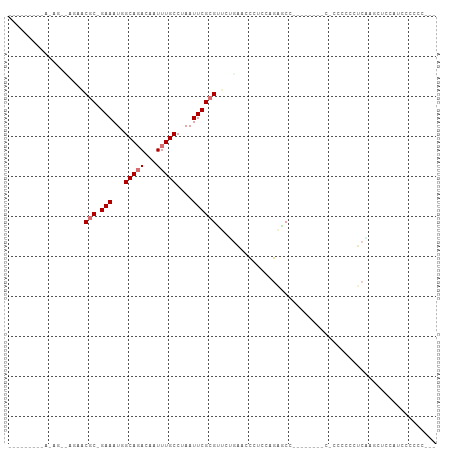
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:56:15 2011