
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 4,133,115 – 4,133,208 |
| Length | 93 |
| Max. P | 0.601302 |

| Location | 4,133,115 – 4,133,208 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 70.33 |
| Shannon entropy | 0.60590 |
| G+C content | 0.54340 |
| Mean single sequence MFE | -29.68 |
| Consensus MFE | -14.58 |
| Energy contribution | -15.21 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.37 |
| Mean z-score | -0.90 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.601302 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 4133115 93 - 23011544 GCAGUG-CAUUUUUAGAACCACUGACAGGGGCAGAAAACUGCC-GUUCGCCGCCAGUUGUUGCUACGGUAAAUAUGGGUGGUAGCCACCCCCAUC .(((((-............)))))...(((((((....)))))-(((.((((((.....((((....)))).....))))))))).....))... ( -30.60, z-score = -1.03, R) >droSim1.chr2L 4088760 91 - 22036055 GCAGUG-UAUUUUUGAAACCACUGACAGGGGCAGAAAACUGCC-GUUCGCCGCCAGUUGUUGCUACGGUAAAUAUGGGUGGUUGCCACCCCUC-- .(((((-............)))))..((((((((....)))))-((..((((((.....((((....)))).....)))))).))....))).-- ( -30.00, z-score = -0.99, R) >droSec1.super_5 2231979 91 - 5866729 GCAGUG-UAUUUUUGAAACCACUGACAGGGGCAGAAAACUGCC-GUUCGCCGCCAGUUGUUGCUACGGUAAAUAUGGGUGCUUGCCACCCCUC-- .(((((-............))))).....(((((....)))))-(((..(((..(((....))).)))..)))..(((((.....)))))...-- ( -26.30, z-score = -0.07, R) >droYak2.chr2L 4155004 91 - 22324452 GCAGUG-UAUUUUUGGAACCACUGACGGGGCCAGAAAACUGCC-GUUCGCCGCCAGUUGUUGCUACGGUAAACAUGGGUGGCAGCCACCCCUC-- (((((.-..(((((((..((......))..)))))))))))).-(((..(((..(((....))).)))..)))..((((((...))))))...-- ( -35.70, z-score = -1.85, R) >droEre2.scaffold_4929 4196356 91 - 26641161 GCAGUG-UACAUCUGGAACCACUGACAGGGGCAGAAAAUUGCC-AUUCGCCGCCAGUUGUUGCUACGGUAAAUAUGGGUGGCAGCCACCCUUC-- .(((((-............)))))...(((((.(((.......-.)))((((((.....((((....)))).....)))))).))).))....-- ( -29.80, z-score = -0.69, R) >droAna3.scaffold_12943 702114 92 + 5039921 ACAGUA-CAGUUUGACAUUCAAUGGCUAUGGCAGAAAAGUGCCUAUCCGAUGGCUCUUGUUGCUACGGUAAAUAUGGGCGUCUGCCACCCCUC-- ..(((.-....(((.....)))..))).(((((((....((((((((((.((((.......))))))).....))))))))))))))......-- ( -26.90, z-score = -1.02, R) >dp4.chr4_group3 9897762 85 + 11692001 GUGGUGCCACUGCCUCUGCCACUGUCACUGGCAGAAAAGUGCC-AGUCGUUGCCAGUUGUUGCUACGGUAAAUAUGGGCGGCCAUC--------- .(((...((((...(((((((.......)))))))..))))))-)...((((((.....((((....)))).....))))))....--------- ( -30.50, z-score = -0.92, R) >droPer1.super_1 7017543 85 + 10282868 GUGGUGCCACUGCCUCUGCCACUGUCACUGGCAGAAAAGUGCC-AGUCGUUGCCAGUUGUUGCUACGGUAAAUAUGGGCGGCCAUC--------- .(((...((((...(((((((.......)))))))..))))))-)...((((((.....((((....)))).....))))))....--------- ( -30.50, z-score = -0.92, R) >droGri2.scaffold_15252 12723324 79 - 17193109 GCAAU--UCAGGCGCCUGUCAAAGGCAGCGGCUGAAAAGUGCC----CGUUGUCAGCUGC--GUACGGCAAACGAGACAUGUUGACG-------- (((.(--((((.(((.(((.....)))))).)))))...))).----....(((((((((--...((.....)).).)).)))))).-------- ( -26.80, z-score = -0.59, R) >consensus GCAGUG_UAUUUUUGCAACCACUGACAGGGGCAGAAAACUGCC_AUUCGCCGCCAGUUGUUGCUACGGUAAAUAUGGGUGGCUGCCACCCCUC__ .(((((.............))))).....((((......)))).....((((((.....((((....)))).....))))))............. (-14.58 = -15.21 + 0.63)
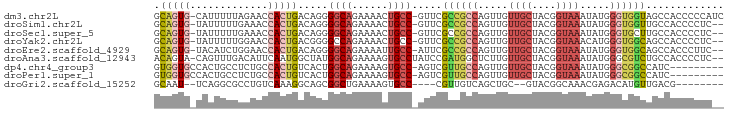


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:15:27 2011