
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 976,084 – 976,188 |
| Length | 104 |
| Max. P | 0.609126 |

| Location | 976,084 – 976,188 |
|---|---|
| Length | 104 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 78.26 |
| Shannon entropy | 0.41387 |
| G+C content | 0.36913 |
| Mean single sequence MFE | -14.63 |
| Consensus MFE | -10.13 |
| Energy contribution | -9.90 |
| Covariance contribution | -0.23 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.609126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 976084 104 + 24543557 --CAUCG--AAAACAACAAAAGCCACACGCAAAGUGGCCUGGUAUUCACACCACUCAAUGAGUAUGCAAUU---------ACAACCAACUAA-CUUCAAUUACGAAAUCUAAAUAUUU --..(((--............(((((.......))))).((((.........((((...)))).((.....---------.)))))).....-.........)))............. ( -13.80, z-score = -1.26, R) >droSim1.chr3L 607842 104 + 22553184 --CAUCG--AAAACAACAAAAGCCACACGCAAAGUGGCCUGGUAUUCACACCACUCAAUGAGUAUGCAAUU---------ACAACCAACUAA-CUUCAAUUACGAAAUCUAAAUAUUU --..(((--............(((((.......))))).((((.........((((...)))).((.....---------.)))))).....-.........)))............. ( -13.80, z-score = -1.26, R) >droSec1.super_2 988040 104 + 7591821 --CAUCG--AAAACAACAAAAGCCACACGCAAAGUGGCCUGGUAUUCACACCACUCAAUGAGUAUGCAAUU---------ACAACCAACUAA-CUUCAAUUACGAAAUCUAAAUAUUU --..(((--............(((((.......))))).((((.........((((...)))).((.....---------.)))))).....-.........)))............. ( -13.80, z-score = -1.26, R) >droEre2.scaffold_4784 976903 104 + 25762168 --CAUCG--AAAACAACAAAAGCCACACGCAAAGUGGCCUGGUAUUCACACCACUCAAUGAGUAUGCAAUU---------ACAACCAACUAA-CUUCAAUUACGAAAUCUAAAUAUUU --..(((--............(((((.......))))).((((.........((((...)))).((.....---------.)))))).....-.........)))............. ( -13.80, z-score = -1.26, R) >droAna3.scaffold_13337 16352028 107 - 23293914 AUCACCGUUACAACAACAAUAGCCACACGCAAAGUGGCCUGGUAUUCACGCCACUCAAUGAGUAUGCAAUU---------ACAACCAACUAA-CUUCAAUUACGAA-UCUAAAUAUUU ......((((...........(((((.......))))).((((......((.((((...))))..))....---------...))))..)))-)............-........... ( -14.42, z-score = -0.66, R) >dp4.chrXR_group6 6592713 103 + 13314419 -------------CGAUG-CAGCCACUCGCAAAGUGGUUUGGUAUUCACGCCACUCAAUGAGUAUGCAAUUUAGUUCCAAAAAACCAAAUAAACUUCAAUUA-AAUAUCGAAAUAUAU -------------(((((-((...(((((...((((((.((.....)).))))))...))))).))).((((((((.....................)))))-)))))))........ ( -20.30, z-score = -2.27, R) >droVir3.scaffold_13049 15535194 99 + 25233164 -ACAACAA-CAAAUUACAGAAGCCACACCCAAAGUGGUUUGGUAUUCACGCCACUCAAUGAGUAUGCAAUU------------UCCAUUUCC-AUGCAACUACAAA--GAAACUAU-- -.......-..........(((((((.......)))))))(((......)))..((..(((((.((((...------------.........-.))))))).))..--))......-- ( -14.22, z-score = -0.43, R) >droGri2.scaffold_15110 17301466 96 + 24565398 ---AACAA-CUAACUAUAGAAGCCAAACCCAAAGCGGUCUGGUAUUUACGCCACUCAAUGAGUAUGCAAUU------------UCCAUAUGC--UACUACUACAAA--CAAACUAU-- ---.....-.......((((.((((.(((......))).)))).))))............((((.(((...------------......)))--))))........--........-- ( -12.90, z-score = -0.52, R) >consensus __CAUCG__AAAACAACAAAAGCCACACGCAAAGUGGCCUGGUAUUCACACCACUCAAUGAGUAUGCAAUU_________ACAACCAACUAA_CUUCAAUUACGAAAUCUAAAUAUUU .....................(((((.......)))))..(((......)))((((...))))....................................................... (-10.13 = -9.90 + -0.23)

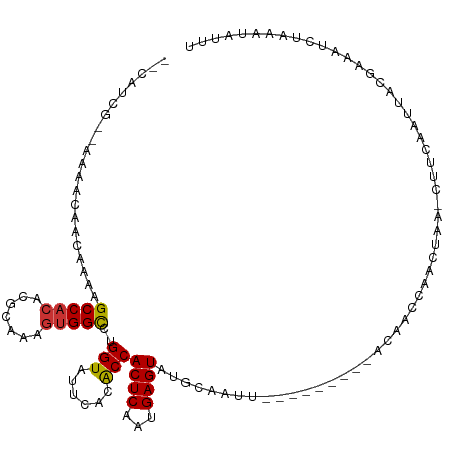

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:54:48 2011