
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 823,330 – 823,424 |
| Length | 94 |
| Max. P | 0.814959 |

| Location | 823,330 – 823,424 |
|---|---|
| Length | 94 |
| Sequences | 10 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 77.84 |
| Shannon entropy | 0.45386 |
| G+C content | 0.40464 |
| Mean single sequence MFE | -22.76 |
| Consensus MFE | -12.01 |
| Energy contribution | -12.05 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.814959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
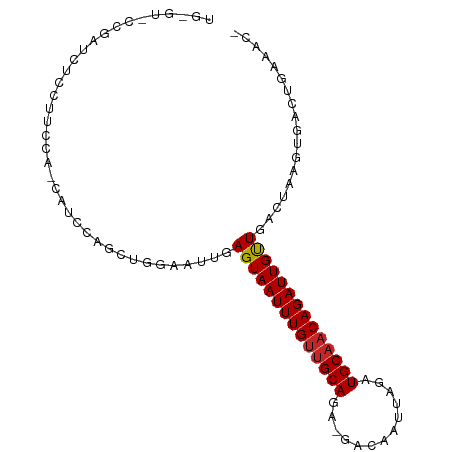
>dm3.chr3L 823330 94 + 24543557 UGAGUGCCGAUCUCCUUUUAGCAUUCAACUGGAAUUGAGUAAUUUGUUGCAGA-GACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGGAAC- (((((((..(.......)..))))))).(..(.(((.((((((((((((((..-..........)))))))))))....))).)))..)..)...- ( -23.50, z-score = -1.62, R) >droSec1.super_2 851167 94 + 7591821 UGAGUGCCGAUCUCCUUUCAGCAUUCAACUGGAAUUGAGUAAUUUGUUGCAGA-GACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGGAAC- (((((((.((.......)).))))))).(..(.(((.((((((((((((((..-..........)))))))))))....))).)))..)..)...- ( -24.60, z-score = -1.56, R) >droYak2.chr3L 818833 94 + 24197627 UGAGGCCCGAUCUGCUUUCAGCAUUCAACUGGAAUUGAGUAAUUUGUUGCAGA-GACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGGAAC- ......(((.((.(((((((((((((((......))))))(((((((((((..-..........))))))))))))))))..)))))).)))...- ( -24.90, z-score = -1.70, R) >droEre2.scaffold_4784 838894 94 + 25762168 UGAGUACCGAUCUCCUUUCAGCAUUCAGCUGGAAUUGAGUAAUUUGUUGCAGA-GACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGGAAC- ......(((.((.(.(((((((.....)))))))(..(.((((((((((((..-..........)))))))))))))..)....).)).)))...- ( -24.00, z-score = -1.40, R) >droAna3.scaffold_13337 11790054 93 + 23293914 UGGGUCCCAGUCUGAGUCUC-AAACUGGACUGAAUUGAGUAAUUUGUUGCAAA-GACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGAAAC- (.(((((((((.((.....)-).)))))(((...(..(.((((((((((((..-..........)))))))))))))..)...))))))).)...- ( -26.00, z-score = -2.23, R) >dp4.chrXR_group8 4803934 79 - 9212921 ---------------UUGCCACAUCUGGCUGGAAUUGAGUAAUUUGUUGCAGA-GACAAUUAGAUGCAACAGAUUGCUGACUAAGUGACUGAAAC- ---------------..((((....))))........((((((((((((((..-..........)))))))))))))).................- ( -21.90, z-score = -2.74, R) >droPer1.super_29 455687 79 - 1099123 ---------------UUGCCACAUCUGGCUGGAAUUGAGUAAUUUGUUGCAGA-GACAAUUAGAUGCAACAGAUUGCUGACUAAGUGACUGAAAC- ---------------..((((....))))........((((((((((((((..-..........)))))))))))))).................- ( -21.90, z-score = -2.74, R) >droWil1.scaffold_180698 9059492 90 + 11422946 UCGGUCUCGAUCUCAGGC----AUCAGCAUGGAAUUGAGUAAUUUGUUGCAAA-GACAAUUAGAUGCUACAGAUUGUUGACUAAGUGACUGAAAC- (((((((((((..((.((----....)).))..))))).((((((((.(((..-..........))).))))))))..........))))))...- ( -22.90, z-score = -1.25, R) >droVir3.scaffold_13049 16438943 83 - 25233164 ----------GCAGCAUCU--CGGCAACACGGAAUUGAGUAAUUUGUUGCAAAAGACAAUUAGAUGCAACAGAUUGUUGACUAAGCGACUGAAAC- ----------.(((...((--..(((((((........))(((((((((((.............))))))))))))))).)..))...)))....- ( -18.52, z-score = -1.19, R) >droMoj3.scaffold_6680 6548459 85 + 24764193 ----------GCAGCAUCAG-CAGCAACACGGAAUUGAGUAAUUUGCUGCAAAAGACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGGAAAC ----------.((((((..(-((((((.((........))...))))))).........((((...((((.....)))).))))))).)))..... ( -19.40, z-score = -1.12, R) >consensus UG_GU_CCGAUCUCCUUCCA_CAUCCAGCUGGAAUUGAGUAAUUUGUUGCAGA_GACAAUUAGAUGCAACAGAUUGUUGACUAAGUGACUGAAAC_ .....................................((((((((((((((.............)))))))))))))).................. (-12.01 = -12.05 + 0.04)


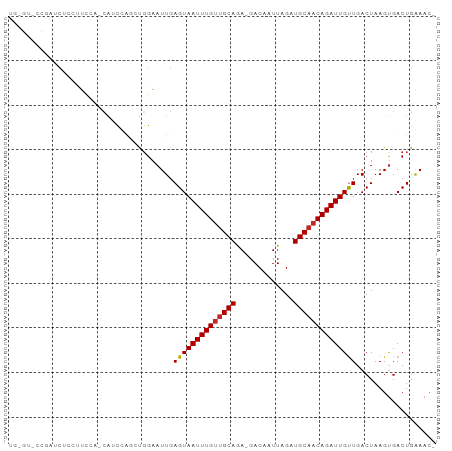
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:54:26 2011