
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 769,550 – 769,650 |
| Length | 100 |
| Max. P | 0.637652 |

| Location | 769,550 – 769,650 |
|---|---|
| Length | 100 |
| Sequences | 10 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 74.10 |
| Shannon entropy | 0.52304 |
| G+C content | 0.45005 |
| Mean single sequence MFE | -17.92 |
| Consensus MFE | -10.89 |
| Energy contribution | -10.71 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.08 |
| Mean z-score | -0.86 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637652 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 769550 100 - 24543557 AAACUUAUUAAAAAAAAACUGGACCAAAAAAUAUAUAGUCACCA--ACGAC-------UUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACACA ..................(((((.((..........((((....--..)))-------))).)))))(((((.......)))))......................... ( -16.60, z-score = -1.59, R) >droGri2.scaffold_15110 19954628 80 + 24565398 --------------AAAAAU------AUGAAAAAAUUGUCGACA--AAGAC-------CUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACAGA --------------.....(------((((......(((((...--...((-------(((...)))))(((.......))).......)))))....)))))...... ( -15.60, z-score = -1.04, R) >droVir3.scaffold_13049 16377945 80 + 25233164 --------------AAAAAU------AAAAAAGAACAGUCGACA--ACGAC-------CUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUUGCUCAUACACAGU --------------......------......((.((((((...--.))))-------.((((....(((((.......)))))......)))).)).))......... ( -14.70, z-score = -0.63, R) >dp4.chrXR_group8 4743938 94 + 9212921 ------AAAUGAAGAAAAGGAGGGCAAAAAAAAUUAAGUCUCAA--CCGAC-------CUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUUGCUCAUACACACA ------...............((((((..........(((....--..)))-------.((((....(((((.......)))))......))))))))))......... ( -19.50, z-score = -0.23, R) >droPer1.super_29 395236 93 + 1099123 ------AAAUGAAGAAAAGGAGGGC-AAAAAAAUUAAGUCUCAA--CCGAC-------CUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUUGCUCAUACACACA ------...............((((-((.........(((....--..)))-------.((((....(((((.......)))))......))))))))))......... ( -19.50, z-score = -0.26, R) >droAna3.scaffold_13337 11722515 93 - 23293914 ------UAAAAUAUAAAAGUUUAGUAAAAA--AUUAAGUCACCA-UGCGAC-------UUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACACA ------............((..(((.....--...(((((....-...)))-------))(((....(((((.......)))))......)))...)))...))..... ( -14.60, z-score = -0.28, R) >droEre2.scaffold_4784 782583 93 - 25762168 ------AAAAAAAAAAAACUGCACCAAAAAG-AUAUAGUCACCA--GCGAC-------UUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACACA ------..............((.........-....((((....--..)))-------)((((....(((((.......)))))......))))..))........... ( -14.60, z-score = -0.38, R) >droYak2.chr3L 764080 93 - 24197627 ------AGAAAAAAAAAACUGGACCAAAAAG-AUAUAGUCACCA--GCGAC-------UUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACACA ------.((.........(((((.((.....-....((((....--..)))-------))).)))))(((((.......)))))..............))......... ( -16.80, z-score = -0.58, R) >droSec1.super_2 789053 94 - 7591821 ------AAAAAAAAAAAACUGUAUCAAAAAAUACAUAGUCACCA--GCGAC-------UUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACACA ------.............(((((......)))))..((....(--((...-------.((((....(((((.......)))))......))))..)))...))..... ( -15.10, z-score = -1.14, R) >anoGam1.chr2L 13947484 105 + 48795086 ---UAAUGUACUGAGUGUGUCUACUAAAAAAGCUUCGGUUAAAGCAACAACACUGUUGCUGCUCCAGGUGAGGCUCGAACUCACAACCACGGCAUCGCUCGAAGCAAC- ---...(((..((((((((((...............((....(((((((....)))))))...))..(((((.......)))))......)))).))))))..)))..- ( -32.20, z-score = -2.43, R) >consensus ______AAAAAAAAAAAACUGGACCAAAAAAAAUAUAGUCACCA__ACGAC_______CUGCUCCAGGUGAGGCUCGAACUCACAACCCCGGCAUAGCUCAUACACACA .....................................(((........)))........((((....(((((.......)))))......))))............... (-10.89 = -10.71 + -0.18)


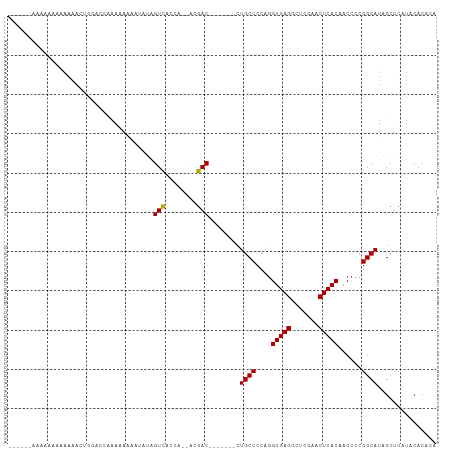
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:54:09 2011