
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 755,832 – 755,896 |
| Length | 64 |
| Max. P | 0.539048 |

| Location | 755,832 – 755,896 |
|---|---|
| Length | 64 |
| Sequences | 12 |
| Columns | 67 |
| Reading direction | reverse |
| Mean pairwise identity | 89.39 |
| Shannon entropy | 0.22602 |
| G+C content | 0.38057 |
| Mean single sequence MFE | -14.17 |
| Consensus MFE | -10.05 |
| Energy contribution | -9.65 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.539048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 755832 64 - 24543557 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCUCGCCAA-AGUCGUCG ...((((((..((((.....))))...))))))((((((...(((..--...))).))-)))).... ( -13.90, z-score = -1.65, R) >droSim1.chr3L 406269 64 - 22553184 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCUCGCCAA-AGUCGUCG ...((((((..((((.....))))...))))))((((((...(((..--...))).))-)))).... ( -13.90, z-score = -1.65, R) >droSec1.super_2 775609 64 - 7591821 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCUCGCCAA-AGUCGUCG ...((((((..((((.....))))...))))))((((((...(((..--...))).))-)))).... ( -13.90, z-score = -1.65, R) >droYak2.chr3L 749719 64 - 24197627 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCUCGCCAA-AGUCGUCG ...((((((..((((.....))))...))))))((((((...(((..--...))).))-)))).... ( -13.90, z-score = -1.65, R) >droEre2.scaffold_4784 768428 59 - 25762168 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCUCGCCAAAGU------ .........((((((((((((.((......)).))))))))))))..--............------ ( -12.40, z-score = -1.69, R) >droAna3.scaffold_13337 11705631 64 - 23293914 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCUCGCCAA-AGUCGUCG ...((((((..((((.....))))...))))))((((((...(((..--...))).))-)))).... ( -13.90, z-score = -1.65, R) >dp4.chrXR_group8 4730176 67 + 9212921 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCACCCCUCGCCAAGAGCCGGCA .........((((((((((((.((......)).))))))))))))........(((.......))). ( -16.70, z-score = -1.84, R) >droPer1.super_29 381254 67 + 1099123 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCACCCCUCGCCAAGAGCCGGCA .........((((((((((((.((......)).))))))))))))........(((.......))). ( -16.70, z-score = -1.84, R) >droWil1.scaffold_180698 8959452 58 - 11422946 AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA--CCU--ACAA-AGUC---- .(((((...((((((((((((.((......)).))))))))))))..--..)--))))-....---- ( -15.60, z-score = -3.55, R) >droVir3.scaffold_13049 16362580 62 + 25233164 UUUGUGUUUCGUGUGAAAAUCACAAUUAAAUGCAAUUUUUAUGCGCA--CCU--ACAA-AACCGUCG ((((((...(((((((((((..((......))..)))))))))))..--..)--))))-)....... ( -11.70, z-score = -1.42, R) >droMoj3.scaffold_6680 6470706 62 - 24764193 UUUGUGUUUCGUGUGAAAAUCACAAUUAAAUGCAAUUUUUAUGCGCA--CCU--ACAA-AGACGUCG ((((((...(((((((((((..((......))..)))))))))))..--..)--))))-)....... ( -11.70, z-score = -0.88, R) >droGri2.scaffold_15110 19936482 62 + 24565398 UUUGUAUUUCGUGUGAAAAUCGCAAUUAAAUGCAAUUUUUAUGCGCA--CCU--ACAA-AGUCGACG ((((((...(((((((((((.(((......))).)))))))))))..--..)--))))-)....... ( -15.70, z-score = -2.54, R) >consensus AUUGUAUUUCGUGUGAAAAUCACAAUUAAAUGCGAUUUUUAUGCGCA__CCUCGCCAA_AGUCGUCG .((((((((..((((.....))))...))))))))................................ (-10.05 = -9.65 + -0.40)

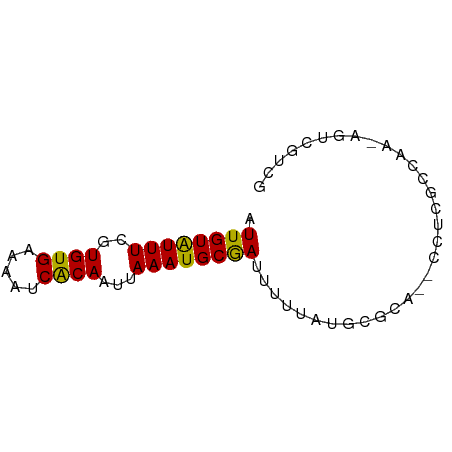
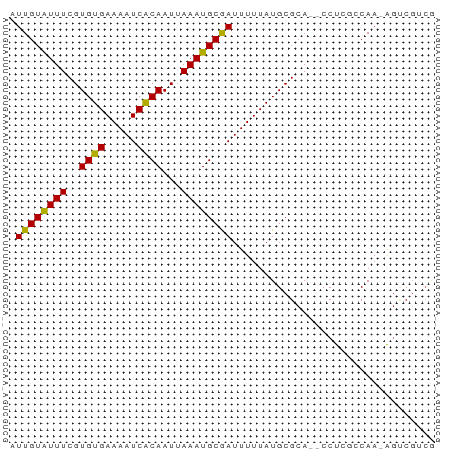
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:54:04 2011