
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 676,508 – 676,609 |
| Length | 101 |
| Max. P | 0.538356 |

| Location | 676,508 – 676,609 |
|---|---|
| Length | 101 |
| Sequences | 9 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 85.81 |
| Shannon entropy | 0.29322 |
| G+C content | 0.40707 |
| Mean single sequence MFE | -24.08 |
| Consensus MFE | -12.49 |
| Energy contribution | -13.01 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.538356 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
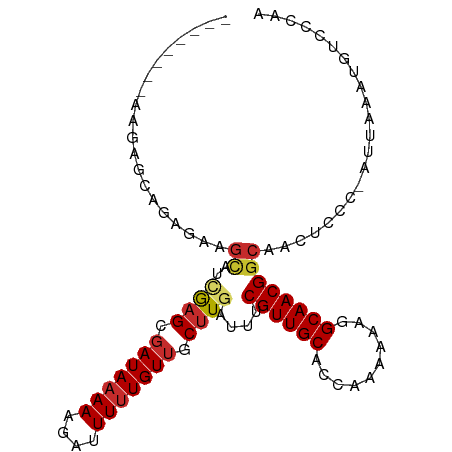
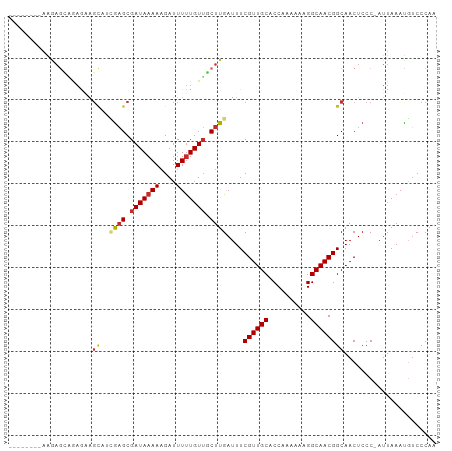
>dm3.chr3L 676508 101 + 24543557 AAAAGCAGAGAAGCAGAGAAGCAUCGAGCGAUAAAAAGAUUUUUGUUGCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAAUGUCCCAA ....(((...((...(((..((((((((((((((((....))))))))))))))..((((((..........))))))))..)))..-.))...)))..... ( -28.50, z-score = -3.24, R) >droSim1.chr3L 325749 93 + 22553184 --------AAGAGCAGAGAAGCAUCGAGCGAUAAAAAGAUUUUUGUUGCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAAUGUCCCAA --------.......(((..((((((((((((((((....))))))))))))))..((((((..........))))))))..)))..-.............. ( -26.90, z-score = -3.26, R) >droSec1.super_2 696642 93 + 7591821 --------AAGAACUGAGAAGCAUCGAGCGAUAAAAAGAUUUUUGUUGCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAAUGUCCCAA --------.......(((..((((((((((((((((....))))))))))))))..((((((..........))))))))..)))..-.............. ( -26.70, z-score = -3.38, R) >droYak2.chr3L 665146 93 + 24197627 --------AAAAGCAGAGAAGCACCGAGCGAUAAAAUGAUUUUUGUUGCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAAUGUCCCAA --------.......(((..((..((((((((((((....))))))))))))....((((((..........))))))))..)))..-.............. ( -25.70, z-score = -2.98, R) >droEre2.scaffold_4784 684949 93 + 25762168 --------AAAAGCAGAGAAGCAUCGAGCGAUAAAAAGAUUUUUGUUGCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAAUGUCCCAA --------.......(((..((((((((((((((((....))))))))))))))..((((((..........))))))))..)))..-.............. ( -26.90, z-score = -3.44, R) >droAna3.scaffold_13337 11620044 93 + 23293914 --------AAGAGUUGAGU-GUAGUGAGCGAUAAAAAGAUUUUUGUAGCUUGAUUUCGUUGCACCAAAAAUGGCAACGGCAACUCCCCAUUAAACGUCCCAA --------..((((((...-.....((((.((((((....)))))).))))....(((((((..........)))))))))))))................. ( -20.90, z-score = -1.29, R) >dp4.chrXR_group8 4648164 93 - 9212921 --------AGGAGAGAAAGAGCUGGGAGCGAUAAAAAGAUUUUUGUUUCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAACCUCCCAA --------.((.(((....((.((((((.......((((........))))....(((((((..........)))))))...)))))-)))....))))).. ( -24.00, z-score = -1.40, R) >droPer1.super_29 294171 93 - 1099123 --------AGGAGAGAAAGAGCUGGGAGCGAUAAAAAGAUUUUUGUUUCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAACCUCCCAA --------.((.(((....((.((((((.......((((........))))....(((((((..........)))))))...)))))-)))....))))).. ( -24.00, z-score = -1.40, R) >droWil1.scaffold_180698 8856672 78 + 11422946 -----------------------ACAUGCGAUAAAAAGAUUUCUGUUUCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC-AUUAAAUGGCCCAA -----------------------...(((((....((((........))))....))(((((..........))))).)))....((-((...))))..... ( -13.10, z-score = 0.03, R) >consensus ________AAGAGCAGAGAAGCAUCGAGCGAUAAAAAGAUUUUUGUUGCUUGAUUUCGUUGCACCAAAAAAGGCAACGGCAACUCCC_AUUAAAUGUCCCAA ....................((..((((.(((((((....))))))).))))....((((((..........))))))))...................... (-12.49 = -13.01 + 0.52)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:53:50 2011