
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 608,769 – 608,867 |
| Length | 98 |
| Max. P | 0.985990 |

| Location | 608,769 – 608,867 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 77.14 |
| Shannon entropy | 0.39393 |
| G+C content | 0.50002 |
| Mean single sequence MFE | -34.33 |
| Consensus MFE | -22.35 |
| Energy contribution | -22.19 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.22 |
| SVM RNA-class probability | 0.985990 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
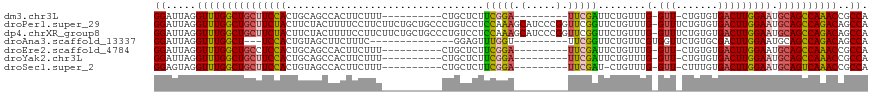
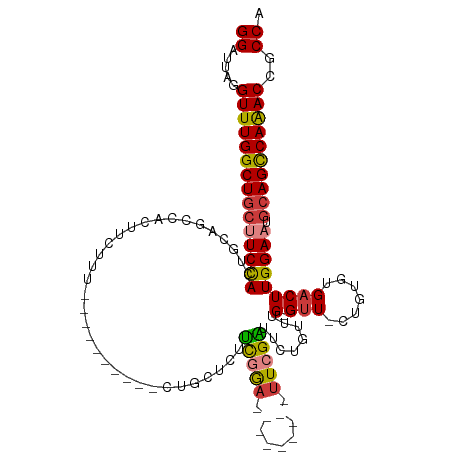
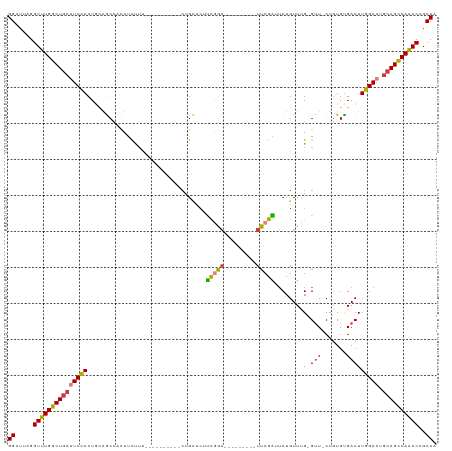
>dm3.chr3L 608769 98 + 24543557 GGAUUAGGUUUGGCUGCUUCCACUGCAGCCACUUCUUU----------CUGCUCUUCGGA---------UUCGAUUCUGUUUG-GUU-CUGUGUGACUUGGAAUGCAGCCAAACCGCCA ((....((((((((((((((((..((((..........----------))))((..((((---------..(((......)))-..)-)))...))..))))).))))))))))).)). ( -38.80, z-score = -4.33, R) >droPer1.super_29 213786 118 - 1099123 GGAUUAGGUUUGGCUGCUUCUACUUCUACUUUUCCUUCUUCUGCUGCCCUGUCCUCCAAAGCAUCCCGGUUCGGUUCUGUUUG-GUUUCUGUGUGACUUGGAAUGCAGCCAGACAGCCA ((.....(((((((((((((((..(((((....((.......((((..(((...............)))..)))).......)-).....))).))..))))).))))))))))..)). ( -33.60, z-score = -1.11, R) >dp4.chrXR_group8 4569679 118 - 9212921 GGAUUAGGUUUGGCUGCUUCUACUUCUACUUUUCCUUCUUCUGCUGCCCUGUCCUCCAAAGCAUCCCGGUUCGGUUCUGUUUG-GUUUCUGUGUGACUUGGAAUGCAGCCAGACAGCCA ((.....(((((((((((((((..(((((....((.......((((..(((...............)))..)))).......)-).....))).))..))))).))))))))))..)). ( -33.60, z-score = -1.11, R) >droAna3.scaffold_13337 16734682 93 - 23293914 GGAUUAGGUUUGGCU---UCCACUGUAGCUUCUUUC--------------GGAGUUUGGU---------UUCGGUUCUGUUCGUGGUUCUGUGCGACUUGGAAUGCAGCCAGACAGCCA ((.....((((((((---.(((....((((((....--------------))))))))).---------....((((((.((((........))))..))))))..))))))))..)). ( -25.80, z-score = 0.15, R) >droEre2.scaffold_4784 612720 98 + 25762168 GGAUUAGGUUUGGCUGCCUCCACUGCAGCCACUUCUUU----------CUGCUCUUCGGA---------UUCGAUUCUGUUUG-GUU-CUGUGUGACUUGGAAUGCAGCCAAACCGCCA ((....(((((((((((.((((..((((..........----------))))((..((((---------..(((......)))-..)-)))...))..))))..))))))))))).)). ( -36.60, z-score = -3.47, R) >droYak2.chr3L 591903 98 + 24197627 GGAUUAGGUUUGGCUGCUUCCACUGCAGCCACUUCUUU----------CUGCUCUUCGGA---------UUCGAUUCUGUUUG-GUU-CUGUGUGACUUGGAAUGCAGCCAAACCGCCA ((....((((((((((((((((..((((..........----------))))((..((((---------..(((......)))-..)-)))...))..))))).))))))))))).)). ( -38.80, z-score = -4.33, R) >droSec1.super_2 629155 97 + 7591821 GGAGUAGGUUUGGCUGCUUCCACUGUAGCCACUUCUUU----------CUGCUCUUCGGA---------UUCGAU-CUGUUUG-GUU-CUUUGUGACUUGGAAUGCAGUCAAACCGCCA ((....((((((((((((((((....((.(((......----------..(((...((((---------(...))-)))...)-)).-....))).))))))).))))))))))).)). ( -33.12, z-score = -2.84, R) >consensus GGAUUAGGUUUGGCUGCUUCCACUGCAGCCACUUCUUU__________CUGCUCUUCGGA_________UUCGAUUCUGUUUG_GUU_CUGUGUGACUUGGAAUGCAGCCAAACCGCCA ((.....(((((((((((((((.................................(((((.(.....).)))))....(((.............))).))))).))))))))))..)). (-22.35 = -22.19 + -0.16)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:53:35 2011