
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 604,346 – 604,408 |
| Length | 62 |
| Max. P | 0.999318 |

| Location | 604,346 – 604,408 |
|---|---|
| Length | 62 |
| Sequences | 7 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 72.72 |
| Shannon entropy | 0.53413 |
| G+C content | 0.31531 |
| Mean single sequence MFE | -12.59 |
| Consensus MFE | -8.69 |
| Energy contribution | -8.90 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.79 |
| SVM RNA-class probability | 0.999318 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
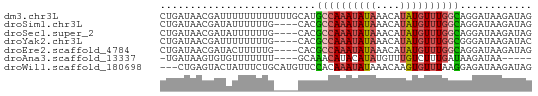
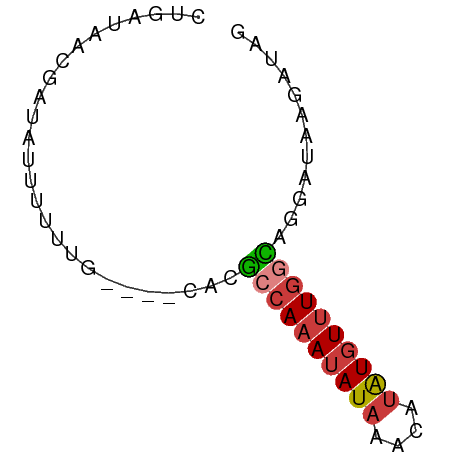
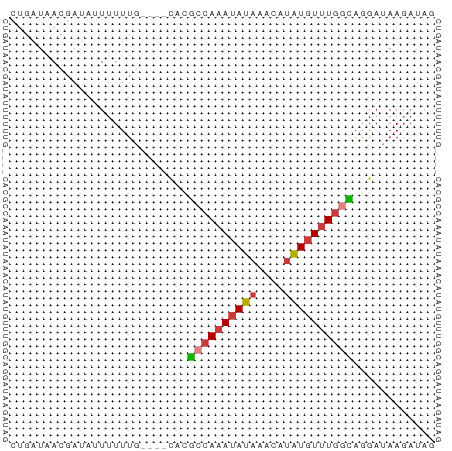
>dm3.chr3L 604346 62 + 24543557 CUGAUAACGAUUUUUUUUUUUUGCAUGCCAAAUAUAAACAUAUGUUUGGCAGGAUAAGAUAG ..................((((((.(((((((((((....))))))))))).).)))))... ( -13.90, z-score = -3.02, R) >droSim1.chr3L 254099 58 + 22553184 CUGAUAACGAUAUUUUUUG----CACGCCAAAUAUAAACAUAUGUUUGGCAGGAUAAGAUAG ..........((((((...----(..((((((((((....))))))))))..)..)))))). ( -14.10, z-score = -3.70, R) >droSec1.super_2 624522 58 + 7591821 CUGAUAACGAUAUUUUUUG----CACGCCAAAUAUAAACAUAUGUUUGGCAGGAUAAGAUAG ..........((((((...----(..((((((((((....))))))))))..)..)))))). ( -14.10, z-score = -3.70, R) >droYak2.chr3L 587201 58 + 24197627 CUGAUAACGAUUUUUUUUG----CACGCCAAAUAUAAACAUAUGUUUGGCGGGAUAAGAUAC ..............(((((----(.(((((((((((....))))))))))).).)))))... ( -14.90, z-score = -3.87, R) >droEre2.scaffold_4784 606856 58 + 25762168 CUGAUAACGAUACUUUUUG----CACGCCAAAUAUAAACAUAUGUUUGGCAGGAUAAGAUAG ..............(((((----..(((((((((((....)))))))))).)..)))))... ( -12.90, z-score = -3.22, R) >droAna3.scaffold_13337 21197318 52 - 23293914 -UGAUAAGUGUGUUUUUUU----GCAAACAUACAUAUGUUUGUCUUUGAUAAGAUAA----- -......((((((((....----..))))))))......(((((((....)))))))----- ( -10.90, z-score = -1.92, R) >droWil1.scaffold_180698 8807952 59 - 11422946 ---CUGAGUACUAUUUCUGCAUGUUCCACAAAUAUAAACAAGUGUUUAAGGAGAUAAGAUAG ---............(((.....((((..((((((......))))))..))))...)))... ( -7.30, z-score = 0.12, R) >consensus CUGAUAACGAUAUUUUUUG____CACGCCAAAUAUAAACAUAUGUUUGGCAGGAUAAGAUAG ..........................((((((((((....))))))))))............ ( -8.69 = -8.90 + 0.21)
| Location | 604,346 – 604,408 |
|---|---|
| Length | 62 |
| Sequences | 7 |
| Columns | 62 |
| Reading direction | reverse |
| Mean pairwise identity | 72.72 |
| Shannon entropy | 0.53413 |
| G+C content | 0.31531 |
| Mean single sequence MFE | -7.96 |
| Consensus MFE | -5.71 |
| Energy contribution | -5.87 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.977638 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
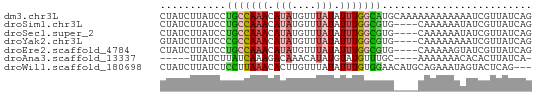
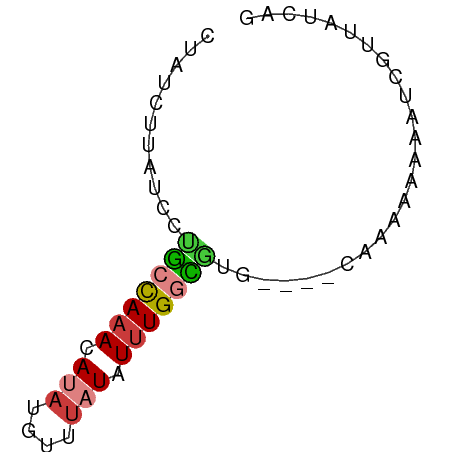
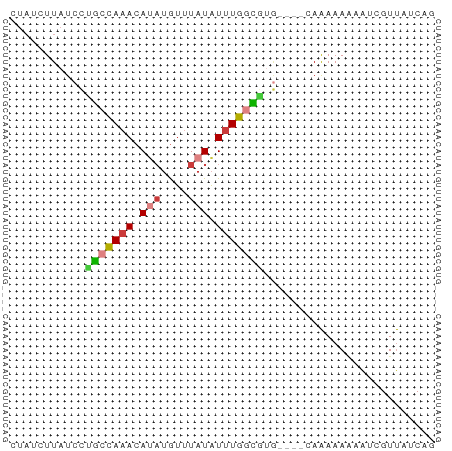
>dm3.chr3L 604346 62 - 24543557 CUAUCUUAUCCUGCCAAACAUAUGUUUAUAUUUGGCAUGCAAAAAAAAAAAAUCGUUAUCAG ...........(((((((.(((....))).)))))))......................... ( -9.10, z-score = -2.20, R) >droSim1.chr3L 254099 58 - 22553184 CUAUCUUAUCCUGCCAAACAUAUGUUUAUAUUUGGCGUG----CAAAAAAUAUCGUUAUCAG ............((((((.(((....))).))))))...----................... ( -8.50, z-score = -1.58, R) >droSec1.super_2 624522 58 - 7591821 CUAUCUUAUCCUGCCAAACAUAUGUUUAUAUUUGGCGUG----CAAAAAAUAUCGUUAUCAG ............((((((.(((....))).))))))...----................... ( -8.50, z-score = -1.58, R) >droYak2.chr3L 587201 58 - 24197627 GUAUCUUAUCCCGCCAAACAUAUGUUUAUAUUUGGCGUG----CAAAAAAAAUCGUUAUCAG ...........(((((((.(((....))).)))))))..----................... ( -10.10, z-score = -2.37, R) >droEre2.scaffold_4784 606856 58 - 25762168 CUAUCUUAUCCUGCCAAACAUAUGUUUAUAUUUGGCGUG----CAAAAAGUAUCGUUAUCAG ............((((((.(((....))).))))))(((----(.....))))......... ( -9.20, z-score = -1.59, R) >droAna3.scaffold_13337 21197318 52 + 23293914 -----UUAUCUUAUCAAAGACAAACAUAUGUAUGUUUGC----AAAAAAACACACUUAUCA- -----...((((....))))(((((((....))))))).----..................- ( -5.70, z-score = -1.41, R) >droWil1.scaffold_180698 8807952 59 + 11422946 CUAUCUUAUCUCCUUAAACACUUGUUUAUAUUUGUGGAACAUGCAGAAAUAGUACUCAG--- ..............(((((....)))))(((((.((.......)).)))))........--- ( -4.60, z-score = 0.82, R) >consensus CUAUCUUAUCCUGCCAAACAUAUGUUUAUAUUUGGCGUG____CAAAAAAAAUCGUUAUCAG ...........(((((((.(((....))).)))))))......................... ( -5.71 = -5.87 + 0.17)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:53:34 2011