
| Sequence ID | dm3.chr3LHet |
|---|---|
| Location | 2,536,892 – 2,536,986 |
| Length | 94 |
| Max. P | 0.999090 |

| Location | 2,536,892 – 2,536,986 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 67.40 |
| Shannon entropy | 0.54227 |
| G+C content | 0.44145 |
| Mean single sequence MFE | -21.47 |
| Consensus MFE | -18.60 |
| Energy contribution | -20.40 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.87 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.27 |
| SVM RNA-class probability | 0.998152 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
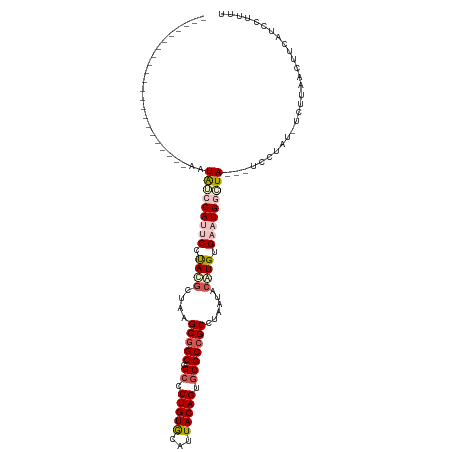
>dm3.chr3LHet 2536892 94 + 2555491 ----------------------AAUAUCCAUUCCUAUGCUAAGCGGCCGCCCUGUGCUUUACAGUGCGCCGCCUAAUACAUGUGAAUGUAUA---UCCUAU-UCUUAAUUACGUACUUUU ----------------------.((((.(((((.((((.((.(((((.((.(((((...))))).))))))).))...)))).))))).)))---).....-.................. ( -23.90, z-score = -3.58, R) >droSec1.super_165 61299 98 + 68140 ----------------------GCUGACCACCCCUACACUAAGCGGCCGCCCUGUAAAUUACAGUGCGCCGCCUAAUACGUGUGAAUGAUUAGAUCGCCAAAAUUCAACUUUAUUAUUUU ----------------------((..................(((((.((.(((((...))))).)))))))(((((.(((....))))))))...))...................... ( -21.60, z-score = -2.54, R) >droSim1.chrU 5059139 90 + 15797150 ----------------------UAUAACCAUCCCUACGCUAAGCAGCCGCUCUGUGCAUUACAGUACGCCGCCUAAUACAUGUGAAUGGUUA---CCCUCUAUCUUUACCCCACC----- ----------------------..(((((((...((((((....))).((..(((((......)))))..)).........))).)))))))---....................----- ( -14.60, z-score = -0.99, R) >droYak2.chrX_random 1718598 107 + 1802292 ---------UGUAAUUUUUCUCAAUAUCCACUCCCAUGCUAAGCGGCCGCCCUGUGCGUUACAGUGCGCCGCCCAAUACAUGUGAAUGGCUA---UCCUGU-UCUUAAUUUUCUCCUUUU ---------..............(((.(((.((.((((....(((((.((.(((((...))))).)))))))......)))).)).))).))---).....-.................. ( -23.80, z-score = -2.49, R) >droEre2.scaffold_4929 26458487 116 + 26641161 UUUAAUUUUUCCUCAAUAUACGAAUAUCCAUUCCUACGCUAAGCGGCCGCCCUGUGCGUUACAGUGCGCCGCCUAAUACAUGUGAAUGGCUA---UCCUAU-UCUUUAAUUAUUUCUUUU .....................(((((.(((((((........(((((.((.(((((...))))).))))))).........).))))))...---...)))-))................ ( -23.43, z-score = -2.23, R) >consensus ______________________AAUAUCCAUUCCUACGCUAAGCGGCCGCCCUGUGCAUUACAGUGCGCCGCCUAAUACAUGUGAAUGGCUA___UCCUAU_UCUUAACUUCAUCCUUUU ........................(((((((((.((((....(((((.((.(((((...))))).)))))))......)))).)))))))))............................ (-18.60 = -20.40 + 1.80)


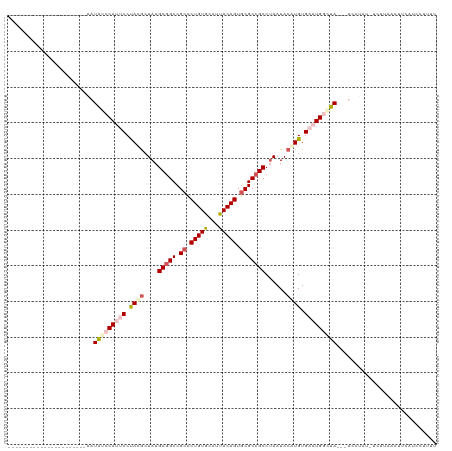
| Location | 2,536,892 – 2,536,986 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 67.40 |
| Shannon entropy | 0.54227 |
| G+C content | 0.44145 |
| Mean single sequence MFE | -32.56 |
| Consensus MFE | -30.06 |
| Energy contribution | -29.06 |
| Covariance contribution | -1.00 |
| Combinations/Pair | 1.37 |
| Mean z-score | -4.05 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.64 |
| SVM RNA-class probability | 0.999090 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
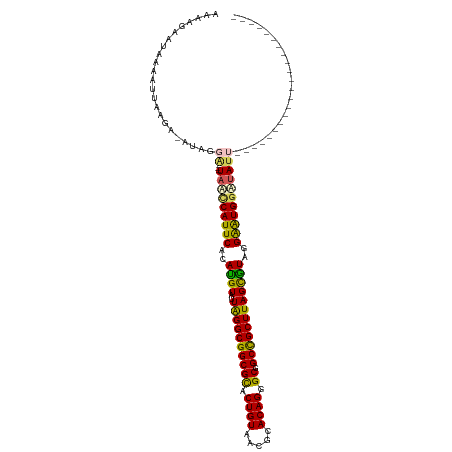
>dm3.chr3LHet 2536892 94 - 2555491 AAAAGUACGUAAUUAAGA-AUAGGA---UAUACAUUCACAUGUAUUAGGCGGCGCACUGUAAAGCACAGGGCGGCCGCUUAGCAUAGGAAUGGAUAUU---------------------- ..................-....((---(((.(((((..((((..((((((((((.((((.....)))).)).))))))))))))..))))).)))))---------------------- ( -30.20, z-score = -4.69, R) >droSec1.super_165 61299 98 - 68140 AAAAUAAUAAAGUUGAAUUUUGGCGAUCUAAUCAUUCACACGUAUUAGGCGGCGCACUGUAAUUUACAGGGCGGCCGCUUAGUGUAGGGGUGGUCAGC---------------------- ...........((..(...)..))...((.(((((((.....(((((((((((((.(((((...))))).)).)))))))))))...))))))).)).---------------------- ( -30.10, z-score = -2.85, R) >droSim1.chrU 5059139 90 - 15797150 -----GGUGGGGUAAAGAUAGAGGG---UAACCAUUCACAUGUAUUAGGCGGCGUACUGUAAUGCACAGAGCGGCUGCUUAGCGUAGGGAUGGUUAUA---------------------- -----...................(---(((((((((..((((..((((((((((.((((.....)))).)).))))))))))))..)))))))))).---------------------- ( -31.60, z-score = -3.60, R) >droYak2.chrX_random 1718598 107 - 1802292 AAAAGGAGAAAAUUAAGA-ACAGGA---UAGCCAUUCACAUGUAUUGGGCGGCGCACUGUAACGCACAGGGCGGCCGCUUAGCAUGGGAGUGGAUAUUGAGAAAAAUUACA--------- ..................-....((---((.((((((.(((((..((((((((((.((((.....)))).)).))))))))))))).)))))).)))).............--------- ( -35.40, z-score = -4.72, R) >droEre2.scaffold_4929 26458487 116 - 26641161 AAAAGAAAUAAUUAAAGA-AUAGGA---UAGCCAUUCACAUGUAUUAGGCGGCGCACUGUAACGCACAGGGCGGCCGCUUAGCGUAGGAAUGGAUAUUCGUAUAUUGAGGAAAAAUUAAA ..................-...(((---((.((((((..((((..((((((((((.((((.....)))).)).))))))))))))..)))))).)))))..................... ( -35.50, z-score = -4.40, R) >consensus AAAAGAAUAAAAUUAAGA_AUAGGA___UAACCAUUCACAUGUAUUAGGCGGCGCACUGUAACGCACAGGGCGGCCGCUUAGCGUAGGAAUGGAUAUU______________________ ............................(((((((((..((((..((((((((((.((((.....)))).)).))))))))))))..)))))))))........................ (-30.06 = -29.06 + -1.00)


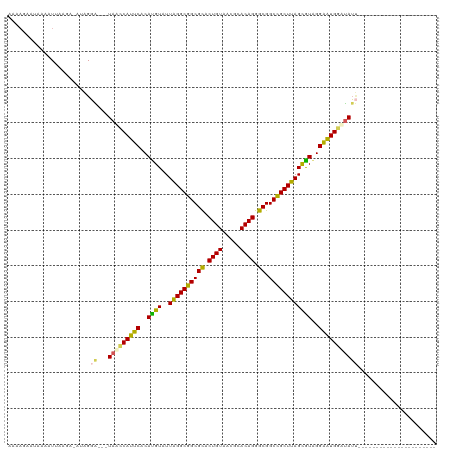
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:52:03 2011